[English] 日本語
 Yorodumi
Yorodumi- EMDB-1574: C-terminal domain of adenovirus serotype 5 protein IX assemble in... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1574 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | C-terminal domain of adenovirus serotype 5 protein IX assemble into an anti-parallel structure | |||||||||
 Map data Map data | SY12 modified adenovirus 5 | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | adenovirus / proteinIX | |||||||||
| Biological species |   Human adenovirus 5 Human adenovirus 5 | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 11.0 Å | |||||||||
 Authors Authors | Fabry CMS / Rosa-Calatrava M / Moriscot C / Ruigrok RWH / Boulanger P / Schoehn G | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2009 Journal: J Virol / Year: 2009Title: The C-terminal domains of adenovirus serotype 5 protein IX assemble into an antiparallel structure on the facets of the capsid. Authors: Céline M S Fabry / Manuel Rosa-Calatrava / Christine Moriscot / Rob W H Ruigrok / Pierre Boulanger / Guy Schoehn /  Abstract: Adenovirus serotype 5 protein IX (pIX) has two domains connected by a flexible linker. Three N-terminal domains form triskelions on the capsid facets that cement hexons together, and the C-terminal ...Adenovirus serotype 5 protein IX (pIX) has two domains connected by a flexible linker. Three N-terminal domains form triskelions on the capsid facets that cement hexons together, and the C-terminal domains of four monomers form complexes toward the facet periphery. Here we present a cryoelectron microscopy structure of recombinant adenovirus with a peptide tag added to the C terminus of pIX. The structure, made up by several C termini of pIX, is longer at both ends than the wild-type protein, and Fabs directed against the tag bind to both ends of the oligomer, demonstrating that the pIX C termini associate in an antiparallel manner. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1574.map.gz emd_1574.map.gz | 72.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1574-v30.xml emd-1574-v30.xml emd-1574.xml emd-1574.xml | 8.8 KB 8.8 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-1574.png EMD-1574.png | 796.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1574 http://ftp.pdbj.org/pub/emdb/structures/EMD-1574 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1574 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1574 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1574.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) Download / File: emd_1574.map.gz / Format: CCP4 / Size: 166.4 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SY12 modified adenovirus 5 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.33 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
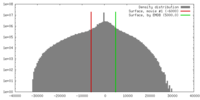
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SY12 modified adenovirus 5
| Entire | Name: SY12 modified adenovirus 5 |
|---|---|
| Components |
|
-Supramolecule #1000: SY12 modified adenovirus 5
| Supramolecule | Name: SY12 modified adenovirus 5 / type: sample / ID: 1000 Details: dodecapeptide TAYSSYMKGGKF (abbreviated SY12 ) fused to the C-terminus of pIX and a recombinant Ad5LacZ-pIX-SY12 vector has been constructed Number unique components: 1 |
|---|
-Supramolecule #1: Human adenovirus 5
| Supramolecule | Name: Human adenovirus 5 / type: virus / ID: 1 / Name.synonym: adenovirus / NCBI-ID: 28285 / Sci species name: Human adenovirus 5 / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No / Syn species name: adenovirus |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Virus shell | Shell ID: 1 / Diameter: 1000 Å / T number (triangulation number): 25 |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1. mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 20mM NaCl, 10mM Tris-HCL |
| Staining | Type: NEGATIVE / Details: Cryo EM |
| Grid | Details: quantifoil r2/2 |
| Vitrification | Cryogen name: ETHANE / Instrument: OTHER / Details: Vitrification instrument: zeiss / Method: Blot for 2 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2010F |
|---|---|
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 100,000 times magnification |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 18 / Average electron dose: 9 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 1.4 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 30000 |
| Sample stage | Specimen holder: Eucentric / Specimen holder model: OTHER |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 11.0 Å / Resolution method: FSC 0.33 CUT-OFF / Software - Name: pft2 em3dr2 / Number images used: 2943 |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)