+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11661 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of VP40 matrix layer in EBOV VP40 VLPs | ||||||||||||
 Map data Map data | Structure of VP40 matrix layer in EBOV VP40 VLPs | ||||||||||||
 Sample Sample |
| ||||||||||||
| Biological species |   | ||||||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 9.8 Å | ||||||||||||
 Authors Authors | Wan W / Clarke M / Norris M / Kolesnikova L / Koehler A / Bornholdt ZA / Becker S / Saphire EO / Briggs JAG | ||||||||||||
| Funding support | European Union,  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Ebola and Marburg virus matrix layers are locally ordered assemblies of VP40 dimers. Authors: William Wan / Mairi Clarke / Michael J Norris / Larissa Kolesnikova / Alexander Koehler / Zachary A Bornholdt / Stephan Becker / Erica Ollmann Saphire / John Ag Briggs /    Abstract: Filoviruses such as Ebola and Marburg virus bud from the host membrane as enveloped virions. This process is achieved by the matrix protein VP40. When expressed alone, VP40 induces budding of ...Filoviruses such as Ebola and Marburg virus bud from the host membrane as enveloped virions. This process is achieved by the matrix protein VP40. When expressed alone, VP40 induces budding of filamentous virus-like particles, suggesting that localization to the plasma membrane, oligomerization into a matrix layer, and generation of membrane curvature are intrinsic properties of VP40. There has been no direct information on the structure of VP40 matrix layers within viruses or virus-like particles. We present structures of Ebola and Marburg VP40 matrix layers in intact virus-like particles, and within intact Marburg viruses. VP40 dimers assemble extended chains via C-terminal domain interactions. These chains stack to form 2D matrix lattices below the membrane surface. These lattices form a patchwork assembly across the membrane and suggesting that assembly may begin at multiple points. Our observations define the structure and arrangement of the matrix protein layer that mediates formation of filovirus particles. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11661.map.gz emd_11661.map.gz | 24.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11661-v30.xml emd-11661-v30.xml emd-11661.xml emd-11661.xml | 11 KB 11 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11661.png emd_11661.png | 79.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11661 http://ftp.pdbj.org/pub/emdb/structures/EMD-11661 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11661 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11661 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11661.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11661.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of VP40 matrix layer in EBOV VP40 VLPs | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.78 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
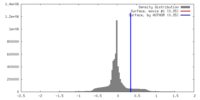
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Ebola virus - Mayinga, Zaire, 1976
| Entire | Name:  |
|---|---|
| Components |
|
-Supramolecule #1: Ebola virus - Mayinga, Zaire, 1976
| Supramolecule | Name: Ebola virus - Mayinga, Zaire, 1976 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 128952 / Sci species name: Ebola virus - Mayinga, Zaire, 1976 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: SPECIES / Virus enveloped: Yes / Virus empty: Yes |
|---|---|
| Host system | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: VP40
| Macromolecule | Name: VP40 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MRRVILPTAP PEYMEAIYPV RSNSTIARGG NSNTGFLTPE SVNGDTPSNP LRPIADDTID HASHTPGSV SSAFILEAMV NVISGPKVLM KQIPIWLPLG VADQKTYSFD STTAAIMLAS Y TITHFGKA TNPLVRVNRL GPGIPDHPLR LLRIGNQAFL QEFVLPPVQL ...String: MRRVILPTAP PEYMEAIYPV RSNSTIARGG NSNTGFLTPE SVNGDTPSNP LRPIADDTID HASHTPGSV SSAFILEAMV NVISGPKVLM KQIPIWLPLG VADQKTYSFD STTAAIMLAS Y TITHFGKA TNPLVRVNRL GPGIPDHPLR LLRIGNQAFL QEFVLPPVQL PQYFTFDLTA LK LITQPLP AATWTDDTPT GSNGALRPGI SFHPKLRPIL LPNKSGKKGN SADLTSPEKI QAI MTSLQD FKIVPIDPTK NIMGIEVPET LVHKLTGKKV TSKNGQPIIP VLLPKYIGLD PVAP GDLTM VITQDCDTCH SPASLPAVIE K |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 2.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C2 (2 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 9.8 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: AV3 / Number subtomograms used: 20352 | ||||||
|---|---|---|---|---|---|---|---|
| Extraction | Number tomograms: 39 / Number images used: 480095 | ||||||
| CTF correction | Software:
| ||||||
| Final angle assignment | Type: OTHER / Software: (Name: TOM, AV3) / Details: Constrained Cross Correlation |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)