[English] 日本語
 Yorodumi
Yorodumi- EMDB-11577: Outer Dynein Arm-Shulin complex - full tail region (Tetrahymena t... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11577 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Outer Dynein Arm-Shulin complex - full tail region (Tetrahymena thermophila) | |||||||||
 Map data Map data | Main map of full ODA-Shulin tail (resampled on full-length ODA-Shulin structure using EMDA). Unsharpened map. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
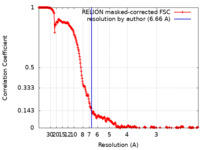
| Method | single particle reconstruction / cryo EM / Resolution: 6.66 Å | |||||||||
 Authors Authors | Mali GR / Abid Ali F / Lau CK / Begum F / Boulanger J / Howe JD / Chen ZA / Rappsilber J / Skehel M / Carter AP | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Shulin packages axonemal outer dynein arms for ciliary targeting. Authors: Girish R Mali / Ferdos Abid Ali / Clinton K Lau / Farida Begum / Jérôme Boulanger / Jonathan D Howe / Zhuo A Chen / Juri Rappsilber / Mark Skehel / Andrew P Carter /   Abstract: The main force generators in eukaryotic cilia and flagella are axonemal outer dynein arms (ODAs). During ciliogenesis, these ~1.8-megadalton complexes are assembled in the cytoplasm and targeted to ...The main force generators in eukaryotic cilia and flagella are axonemal outer dynein arms (ODAs). During ciliogenesis, these ~1.8-megadalton complexes are assembled in the cytoplasm and targeted to cilia by an unknown mechanism. Here, we used the ciliate to identify two factors (Q22YU3 and Q22MS1) that bind ODAs in the cytoplasm and are required for ODA delivery to cilia. Q22YU3, which we named Shulin, locked the ODA motor domains into a closed conformation and inhibited motor activity. Cryo-electron microscopy revealed how Shulin stabilized this compact form of ODAs by binding to the dynein tails. Our findings provide a molecular explanation for how newly assembled dyneins are packaged for delivery to the cilia. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11577.map.gz emd_11577.map.gz | 323 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11577-v30.xml emd-11577-v30.xml emd-11577.xml emd-11577.xml | 16.4 KB 16.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11577_fsc.xml emd_11577_fsc.xml | 27.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_11577.png emd_11577.png | 33.7 KB | ||
| Masks |  emd_11577_msk_1.map emd_11577_msk_1.map | 347.6 MB |  Mask map Mask map | |
| Others |  emd_11577_additional_1.map.gz emd_11577_additional_1.map.gz emd_11577_half_map_1.map.gz emd_11577_half_map_1.map.gz emd_11577_half_map_2.map.gz emd_11577_half_map_2.map.gz | 3.1 MB 1.4 GB 1.4 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11577 http://ftp.pdbj.org/pub/emdb/structures/EMD-11577 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11577 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11577 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11577.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11577.map.gz / Format: CCP4 / Size: 347.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Main map of full ODA-Shulin tail (resampled on full-length ODA-Shulin structure using EMDA). Unsharpened map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11577_msk_1.map emd_11577_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
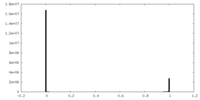
| Density Histograms |
-Additional map: Mask for full ODA-Shulin tail (Z-flipped, non resampled)...
| File | emd_11577_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Mask for full ODA-Shulin tail (Z-flipped, non resampled) to be used with half maps. | ||||||||||||
| Projections & Slices |
| ||||||||||||
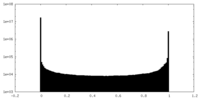
| Density Histograms |
-Half map: Half map 2 of full ODA-Shulin tail (Z-flipped, non-resampled)
| File | emd_11577_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 2 of full ODA-Shulin tail (Z-flipped, non-resampled) | ||||||||||||
| Projections & Slices |
| ||||||||||||
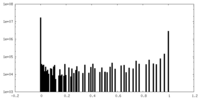
| Density Histograms |
-Half map: Half map 1 of full ODA-Shulin tail (Z-flipped, non-resampled)
| File | emd_11577_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map 1 of full ODA-Shulin tail (Z-flipped, non-resampled) | ||||||||||||
| Projections & Slices |
| ||||||||||||
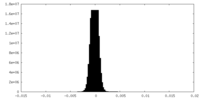
| Density Histograms |
- Sample components
Sample components
-Entire : Tetrahymena thermophila ODA Dyh4 motor
| Entire | Name: Tetrahymena thermophila ODA Dyh4 motor |
|---|---|
| Components |
|
-Supramolecule #1: Tetrahymena thermophila ODA Dyh4 motor
| Supramolecule | Name: Tetrahymena thermophila ODA Dyh4 motor / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.8 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 7.4 |
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Support film - Material: GRAPHENE OXIDE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)