+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-9240 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Rabbit 80S ribosome with eEF2 and SERBP1 (unrotated state with 40S head swivel) | |||||||||
 マップデータ マップデータ | Postprocessed map | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | Ribosome / translation | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報L13a-mediated translational silencing of Ceruloplasmin expression / Translation initiation complex formation / Formation of a pool of free 40S subunits / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Protein hydroxylation ...L13a-mediated translational silencing of Ceruloplasmin expression / Translation initiation complex formation / Formation of a pool of free 40S subunits / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Protein hydroxylation / Translation initiation complex formation / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / L13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / Formation of a pool of free 40S subunits / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Major pathway of rRNA processing in the nucleolus and cytosol / SRP-dependent cotranslational protein targeting to membrane / L13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / Major pathway of rRNA processing in the nucleolus and cytosol / Formation of a pool of free 40S subunits / GTP hydrolysis and joining of the 60S ribosomal subunit / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Translation initiation complex formation / Formation of the ternary complex, and subsequently, the 43S complex / Ribosomal scanning and start codon recognition / Resolution of Sister Chromatid Cohesion / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / RHO GTPases Activate Formins / Major pathway of rRNA processing in the nucleolus and cytosol / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression / SRP-dependent cotranslational protein targeting to membrane / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / Separation of Sister Chromatids / oxidized pyrimidine DNA binding / response to TNF agonist / positive regulation of base-excision repair / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage / negative regulation of DNA repair / supercoiled DNA binding / TORC2 complex binding / NF-kappaB complex / oxidized purine DNA binding / exit from mitosis / ubiquitin-like protein conjugating enzyme binding / Formation of the ternary complex, and subsequently, the 43S complex / erythrocyte homeostasis / optic nerve development / cytoplasmic side of rough endoplasmic reticulum membrane / retinal ganglion cell axon guidance / protein kinase A binding / Ribosomal scanning and start codon recognition / Translation initiation complex formation / positive regulation of T cell receptor signaling pathway / positive regulation of activated T cell proliferation / SARS-CoV-1 modulates host translation machinery / Peptide chain elongation / Selenocysteine synthesis / Formation of a pool of free 40S subunits / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ubiquitin ligase inhibitor activity / Eukaryotic Translation Termination / positive regulation of signal transduction by p53 class mediator / Response of EIF2AK4 (GCN2) to amino acid deficiency / SRP-dependent cotranslational protein targeting to membrane / Viral mRNA Translation / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / 90S preribosome / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression / phagocytic cup / Major pathway of rRNA processing in the nucleolus and cytosol / protein-RNA complex assembly / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / spindle assembly / ribosomal small subunit export from nucleus / rough endoplasmic reticulum / translation regulator activity / translation initiation factor binding / gastrulation / cytosolic ribosome / positive regulation of microtubule polymerization / MDM2/MDM4 family protein binding / negative regulation of protein ubiquitination / positive regulation of interleukin-2 production / class I DNA-(apurinic or apyrimidinic site) endonuclease activity / DNA-(apurinic or apyrimidinic site) lyase / Hsp70 protein binding / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosomal large subunit biogenesis 類似検索 - 分子機能 | |||||||||
| 生物種 |  | |||||||||
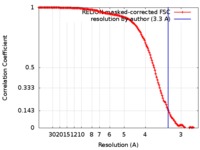
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.3 Å | |||||||||
 データ登録者 データ登録者 | Brown A / Baird MR / Yip MCJ / Murray J / Shao S | |||||||||
 引用 引用 |  ジャーナル: Elife / 年: 2018 ジャーナル: Elife / 年: 2018タイトル: Structures of translationally inactive mammalian ribosomes. 著者: Alan Brown / Matthew R Baird / Matthew Cj Yip / Jason Murray / Sichen Shao /   要旨: The cellular levels and activities of ribosomes directly regulate gene expression during numerous physiological processes. The mechanisms that globally repress translation are incompletely understood. ...The cellular levels and activities of ribosomes directly regulate gene expression during numerous physiological processes. The mechanisms that globally repress translation are incompletely understood. Here, we use electron cryomicroscopy to analyze inactive ribosomes isolated from mammalian reticulocytes, the penultimate stage of red blood cell differentiation. We identify two types of ribosomes that are translationally repressed by protein interactions. The first comprises ribosomes sequestered with elongation factor 2 (eEF2) by SERPINE mRNA binding protein 1 (SERBP1) occupying the ribosomal mRNA entrance channel. The second type are translationally repressed by a novel ribosome-binding protein, interferon-related developmental regulator 2 (IFRD2), which spans the P and E sites and inserts a C-terminal helix into the mRNA exit channel to preclude translation. IFRD2 binds ribosomes with a tRNA occupying a noncanonical binding site, the 'Z site', on the ribosome. These structures provide functional insights into how ribosomal interactions may suppress translation to regulate gene expression. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_9240.map.gz emd_9240.map.gz | 16.7 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-9240-v30.xml emd-9240-v30.xml emd-9240.xml emd-9240.xml | 105 KB 105 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_9240_fsc.xml emd_9240_fsc.xml | 14.1 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_9240.png emd_9240.png | 174 KB | ||
| Filedesc metadata |  emd-9240.cif.gz emd-9240.cif.gz | 21.3 KB | ||
| その他 |  emd_9240_additional.map.gz emd_9240_additional.map.gz emd_9240_half_map_1.map.gz emd_9240_half_map_1.map.gz emd_9240_half_map_2.map.gz emd_9240_half_map_2.map.gz | 214.1 MB 214.4 MB 214.4 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9240 http://ftp.pdbj.org/pub/emdb/structures/EMD-9240 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9240 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9240 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_9240_validation.pdf.gz emd_9240_validation.pdf.gz | 761.3 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_9240_full_validation.pdf.gz emd_9240_full_validation.pdf.gz | 760.9 KB | 表示 | |
| XML形式データ |  emd_9240_validation.xml.gz emd_9240_validation.xml.gz | 22 KB | 表示 | |
| CIF形式データ |  emd_9240_validation.cif.gz emd_9240_validation.cif.gz | 29.1 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9240 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9240 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9240 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9240 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  6mtdMC  9234C  9235C  9236C  9237C  9239C  9241C  9242C  6mtbC  6mtcC  6mteC C: 同じ文献を引用 ( M: このマップから作成された原子モデル |
|---|---|
| 類似構造データ |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_9240.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_9240.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Postprocessed map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
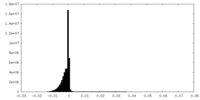
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
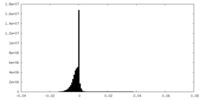
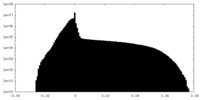
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-追加マップ: Pre-postprocessed map
| ファイル | emd_9240_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Pre-postprocessed map | ||||||||||||
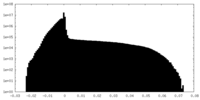
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Half map 1
| ファイル | emd_9240_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 1 | ||||||||||||
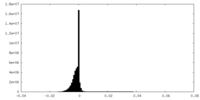
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: Half map 2
| ファイル | emd_9240_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Half map 2 | ||||||||||||
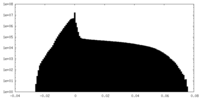
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : Rabbit 80S ribosome with eEF2 and SERBP1 (unrotated state with he...
+超分子 #1: Rabbit 80S ribosome with eEF2 and SERBP1 (unrotated state with he...
+分子 #1: 28S rRNA
+分子 #2: 5S rRNA
+分子 #3: 5.8S rRNA
+分子 #51: 18S rRNA
+分子 #4: uL2
+分子 #5: uL3
+分子 #6: uL4
+分子 #7: uL18
+分子 #8: eL6
+分子 #9: uL30
+分子 #10: eL8
+分子 #11: uL6
+分子 #12: uL16
+分子 #13: uL5
+分子 #14: eL13
+分子 #15: eL14
+分子 #16: eL15
+分子 #17: uL13
+分子 #18: uL22
+分子 #19: eL18
+分子 #20: eL19
+分子 #21: eL20
+分子 #22: eL21
+分子 #23: eL22
+分子 #24: uL14
+分子 #25: eL24
+分子 #26: uL23
+分子 #27: uL24
+分子 #28: eL27
+分子 #29: uL15
+分子 #30: eL29
+分子 #31: eL30
+分子 #32: eL31
+分子 #33: eL32
+分子 #34: eL33
+分子 #35: eL34
+分子 #36: uL29
+分子 #37: eL36
+分子 #38: eL37
+分子 #39: eL38
+分子 #40: eL39
+分子 #41: eL40
+分子 #42: eL41
+分子 #43: eL42
+分子 #44: eL43
+分子 #45: el28
+分子 #46: uL10
+分子 #47: eL11
+分子 #48: uL1
+分子 #49: eEF2
+分子 #50: SERBP1
+分子 #52: uS2
+分子 #53: eS1
+分子 #54: uS5
+分子 #55: uS3
+分子 #56: eS4
+分子 #57: uS7
+分子 #58: eS6
+分子 #59: eS7
+分子 #60: eS8
+分子 #61: uS4
+分子 #62: eS10
+分子 #63: uS17
+分子 #64: eS12
+分子 #65: uS15
+分子 #66: uS11
+分子 #67: uS19
+分子 #68: uS9
+分子 #69: eS17
+分子 #70: uS13
+分子 #71: eS19
+分子 #72: uS10
+分子 #73: eS21
+分子 #74: uS8
+分子 #75: uS12
+分子 #76: eS24
+分子 #77: eS25
+分子 #78: eS26
+分子 #79: eS27
+分子 #80: eS28
+分子 #81: uS14
+分子 #82: eS30
+分子 #83: eS31
+分子 #84: RACK1
+分子 #85: MAGNESIUM ION
+分子 #86: ZINC ION
+分子 #87: GUANOSINE-5'-DIPHOSPHATE
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.4 |
|---|---|
| グリッド | 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: CONTINUOUS / 支持フィルム - Film thickness: 5 |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK II |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: FEI FALCON II (4k x 4k) 検出モード: INTEGRATING / デジタル化 - 画像ごとのフレーム数: 1-17 / 平均露光時間: 1.1 sec. / 平均電子線量: 40.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.7 mm |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー






































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)