[English] 日本語
 Yorodumi
Yorodumi- EMDB-11310: SARS-CoV-2 Nsp1 bound to a pre-40S-like ribosome complex - state 2 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11310 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 Nsp1 bound to a pre-40S-like ribosome complex - state 2 | ||||||||||||
 Map data Map data | SARS-CoV-2 bound to a human pre-40S-like ribosome complex state 2 | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Translational Inhibition / SARS-CoV-2 / Immune Evasion / Human Ribosome / VIRAL PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationendonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / negative regulation of endoplasmic reticulum unfolded protein response / oxidized pyrimidine DNA binding / response to TNF agonist / positive regulation of base-excision repair / positive regulation of ubiquitin-protein transferase activity / positive regulation of respiratory burst involved in inflammatory response / positive regulation of gastrulation / protein tyrosine kinase inhibitor activity / positive regulation of DNA-templated transcription initiation ...endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / negative regulation of endoplasmic reticulum unfolded protein response / oxidized pyrimidine DNA binding / response to TNF agonist / positive regulation of base-excision repair / positive regulation of ubiquitin-protein transferase activity / positive regulation of respiratory burst involved in inflammatory response / positive regulation of gastrulation / protein tyrosine kinase inhibitor activity / positive regulation of DNA-templated transcription initiation / positive regulation of intrinsic apoptotic signaling pathway in response to DNA damage / IRE1-RACK1-PP2A complex / positive regulation of Golgi to plasma membrane protein transport / nucleolus organization / TNFR1-mediated ceramide production / U3 snoRNA binding / negative regulation of RNA splicing / neural crest cell differentiation / supercoiled DNA binding / cytoplasmic translational initiation / NF-kappaB complex / negative regulation of DNA repair / oxidized purine DNA binding / cysteine-type endopeptidase activator activity involved in apoptotic process / rRNA modification in the nucleus and cytosol / negative regulation of intrinsic apoptotic signaling pathway in response to hydrogen peroxide / negative regulation of bicellular tight junction assembly / ubiquitin-like protein conjugating enzyme binding / regulation of establishment of cell polarity / negative regulation of phagocytosis / erythrocyte homeostasis / cytoplasmic side of rough endoplasmic reticulum membrane / Formation of the ternary complex, and subsequently, the 43S complex / ion channel inhibitor activity / protein kinase A binding / laminin receptor activity / Ribosomal scanning and start codon recognition / pigmentation / positive regulation of mitochondrial depolarization / Translation initiation complex formation / negative regulation of Wnt signaling pathway / fibroblast growth factor binding / Protein hydroxylation / monocyte chemotaxis / BH3 domain binding / negative regulation of translational frameshifting / regulation of adenylate cyclase-activating G protein-coupled receptor signaling pathway / SARS-CoV-1 modulates host translation machinery / positive regulation of GTPase activity / TOR signaling / mTORC1-mediated signalling / iron-sulfur cluster binding / Peptide chain elongation / regulation of cell division / cellular response to ethanol / Selenocysteine synthesis / Formation of a pool of free 40S subunits / negative regulation of protein binding / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / Eukaryotic Translation Termination / protein serine/threonine kinase inhibitor activity / SRP-dependent cotranslational protein targeting to membrane / Response of EIF2AK4 (GCN2) to amino acid deficiency / ubiquitin ligase inhibitor activity / negative regulation of respiratory burst involved in inflammatory response / Viral mRNA Translation / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / positive regulation of signal transduction by p53 class mediator / negative regulation of ubiquitin-dependent protein catabolic process / GTP hydrolysis and joining of the 60S ribosomal subunit / L13a-mediated translational silencing of Ceruloplasmin expression / Major pathway of rRNA processing in the nucleolus and cytosol / regulation of translational fidelity / positive regulation of microtubule polymerization / phagocytic cup / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / spindle assembly / positive regulation of intrinsic apoptotic signaling pathway / Protein methylation / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / translation regulator activity / Nuclear events stimulated by ALK signaling in cancer / rough endoplasmic reticulum / ribosomal small subunit export from nucleus / positive regulation of cell cycle / laminin binding / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / translation initiation factor binding / DNA-(apurinic or apyrimidinic site) endonuclease activity / gastrulation / Maturation of protein E / Maturation of protein E / ER Quality Control Compartment (ERQC) / Myoclonic epilepsy of Lafora / signaling adaptor activity / MDM2/MDM4 family protein binding / FLT3 signaling by CBL mutants / negative regulation of protein ubiquitination / IRAK2 mediated activation of TAK1 complex Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.2 Å | ||||||||||||
 Authors Authors | Thoms M / Buschauer R | ||||||||||||
| Funding support |  Germany, 3 items Germany, 3 items
| ||||||||||||
 Citation Citation |  Journal: Science / Year: 2020 Journal: Science / Year: 2020Title: Structural basis for translational shutdown and immune evasion by the Nsp1 protein of SARS-CoV-2. Authors: Matthias Thoms / Robert Buschauer / Michael Ameismeier / Lennart Koepke / Timo Denk / Maximilian Hirschenberger / Hanna Kratzat / Manuel Hayn / Timur Mackens-Kiani / Jingdong Cheng / Jan H ...Authors: Matthias Thoms / Robert Buschauer / Michael Ameismeier / Lennart Koepke / Timo Denk / Maximilian Hirschenberger / Hanna Kratzat / Manuel Hayn / Timur Mackens-Kiani / Jingdong Cheng / Jan H Straub / Christina M Stürzel / Thomas Fröhlich / Otto Berninghausen / Thomas Becker / Frank Kirchhoff / Konstantin M J Sparrer / Roland Beckmann /  Abstract: Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causative agent of the current coronavirus disease 2019 (COVID-19) pandemic. A major virulence factor of SARS-CoVs is the ...Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causative agent of the current coronavirus disease 2019 (COVID-19) pandemic. A major virulence factor of SARS-CoVs is the nonstructural protein 1 (Nsp1), which suppresses host gene expression by ribosome association. Here, we show that Nsp1 from SARS-CoV-2 binds to the 40 ribosomal subunit, resulting in shutdown of messenger RNA (mRNA) translation both in vitro and in cells. Structural analysis by cryo-electron microscopy of in vitro-reconstituted Nsp1-40 and various native Nsp1-40 and -80 complexes revealed that the Nsp1 C terminus binds to and obstructs the mRNA entry tunnel. Thereby, Nsp1 effectively blocks retinoic acid-inducible gene I-dependent innate immune responses that would otherwise facilitate clearance of the infection. Thus, the structural characterization of the inhibitory mechanism of Nsp1 may aid structure-based drug design against SARS-CoV-2. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11310.map.gz emd_11310.map.gz | 106.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11310-v30.xml emd-11310-v30.xml emd-11310.xml emd-11310.xml | 52.7 KB 52.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11310_fsc.xml emd_11310_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_11310.png emd_11310.png | 33.6 KB | ||
| Filedesc metadata |  emd-11310.cif.gz emd-11310.cif.gz | 11.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11310 http://ftp.pdbj.org/pub/emdb/structures/EMD-11310 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11310 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11310 | HTTPS FTP |
-Related structure data
| Related structure data |  6zn5MC  6zlwC  6zm7C  6zmeC  6zmiC  6zmoC  6zmtC  6zonC  6zp4C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11310.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11310.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SARS-CoV-2 bound to a human pre-40S-like ribosome complex state 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.059 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
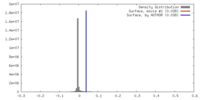
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : SARS-CoV-2 Nsp1 bound to a human pre-40S-like ribosome complex
+Supramolecule #1: SARS-CoV-2 Nsp1 bound to a human pre-40S-like ribosome complex
+Supramolecule #2: human pre-40S-like ribosome complex
+Supramolecule #3: SARS-CoV-2 Nsp1
+Macromolecule #1: 40S ribosomal protein SA
+Macromolecule #2: 40S ribosomal protein S3a
+Macromolecule #3: 40S ribosomal protein S2
+Macromolecule #4: 40S ribosomal protein S4, X isoform
+Macromolecule #5: 40S ribosomal protein S3
+Macromolecule #6: 40S ribosomal protein S6
+Macromolecule #7: 40S ribosomal protein S7
+Macromolecule #8: 40S ribosomal protein S8
+Macromolecule #9: 40S ribosomal protein S9
+Macromolecule #10: 40S ribosomal protein S5
+Macromolecule #11: 40S ribosomal protein S11
+Macromolecule #12: 40S ribosomal protein S10
+Macromolecule #13: 40S ribosomal protein S13
+Macromolecule #14: 40S ribosomal protein S12
+Macromolecule #15: 40S ribosomal protein S14
+Macromolecule #16: 40S ribosomal protein S15
+Macromolecule #17: 40S ribosomal protein S16
+Macromolecule #18: 40S ribosomal protein S17
+Macromolecule #19: 40S ribosomal protein S18
+Macromolecule #20: 40S ribosomal protein S19
+Macromolecule #21: 40S ribosomal protein S20
+Macromolecule #22: 40S ribosomal protein S15a
+Macromolecule #23: 40S ribosomal protein S23
+Macromolecule #24: 40S ribosomal protein S24
+Macromolecule #25: 40S ribosomal protein S21
+Macromolecule #26: 40S ribosomal protein S25
+Macromolecule #27: 40S ribosomal protein S27
+Macromolecule #28: 40S ribosomal protein S26
+Macromolecule #29: 40S ribosomal protein S28
+Macromolecule #30: 40S ribosomal protein S30
+Macromolecule #31: 40S ribosomal protein S29
+Macromolecule #32: Ribosomal protein S27a
+Macromolecule #33: Receptor of activated protein C kinase 1
+Macromolecule #35: Non-structural protein 1
+Macromolecule #36: Pre-rRNA-processing protein TSR1 homolog
+Macromolecule #34: 18S ribosomal RNA
+Macromolecule #37: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 44.8 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)