[English] 日本語
 Yorodumi
Yorodumi- EMDB-11270: E2 core of the fungal Pyruvate dehydrogenase complex with asymmet... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11270 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | E2 core of the fungal Pyruvate dehydrogenase complex with asymmetric interior PX30 component | ||||||||||||||||||
 Map data Map data | Fungal PDC (N. crassa). Recombinant preparation-tE2 PX30. Enforced symmetry: I2. Periphery (PX only) is absent/oversym. Core is structured. Interior is oversymm. | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | acetyl transferase / pyruvate dehydrogenase / protein complex / mitochondria / metabolism / tetrahedral icosahedral / TRANSFERASE | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationdihydrolipoyllysine-residue acetyltransferase / dihydrolipoyllysine-residue acetyltransferase activity / pyruvate decarboxylation to acetyl-CoA / pyruvate dehydrogenase complex / mitochondrial matrix Similarity search - Function | ||||||||||||||||||
| Biological species |  Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) | ||||||||||||||||||
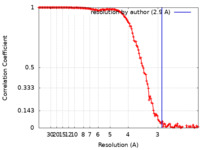
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | ||||||||||||||||||
 Authors Authors | Forsberg BO / Howard RJ / Aibara S / Mortesaei N / Lindahl E | ||||||||||||||||||
| Funding support |  Sweden, European Union, 5 items Sweden, European Union, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Arrangement and symmetry of the fungal E3BP-containing core of the pyruvate dehydrogenase complex. Authors: B O Forsberg / S Aibara / R J Howard / N Mortezaei / E Lindahl /   Abstract: The pyruvate dehydrogenase complex (PDC) is a multienzyme complex central to aerobic respiration, connecting glycolysis to mitochondrial oxidation of pyruvate. Similar to the E3-binding protein (E3BP) ...The pyruvate dehydrogenase complex (PDC) is a multienzyme complex central to aerobic respiration, connecting glycolysis to mitochondrial oxidation of pyruvate. Similar to the E3-binding protein (E3BP) of mammalian PDC, PX selectively recruits E3 to the fungal PDC, but its divergent sequence suggests a distinct structural mechanism. Here, we report reconstructions of PDC from the filamentous fungus Neurospora crassa by cryo-electron microscopy, where we find protein X (PX) interior to the PDC core as opposed to substituting E2 core subunits as in mammals. Steric occlusion limits PX binding, resulting in predominantly tetrahedral symmetry, explaining previous observations in Saccharomyces cerevisiae. The PX-binding site is conserved in (and specific to) fungi, and complements possible C-terminal binding motifs in PX that are absent in mammalian E3BP. Consideration of multiple symmetries thus reveals a differential structural basis for E3BP-like function in fungal PDC. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11270.map.gz emd_11270.map.gz | 36.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11270-v30.xml emd-11270-v30.xml emd-11270.xml emd-11270.xml | 16.7 KB 16.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11270_fsc.xml emd_11270_fsc.xml | 15.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_11270.png emd_11270.png | 64 KB | ||
| Filedesc metadata |  emd-11270.cif.gz emd-11270.cif.gz | 5.6 KB | ||
| Others |  emd_11270_half_map_1.map.gz emd_11270_half_map_1.map.gz emd_11270_half_map_2.map.gz emd_11270_half_map_2.map.gz | 171 MB 171 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11270 http://ftp.pdbj.org/pub/emdb/structures/EMD-11270 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11270 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11270 | HTTPS FTP |
-Related structure data
| Related structure data |  6zloMC  6zlmC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11270.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11270.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Fungal PDC (N. crassa). Recombinant preparation-tE2 PX30. Enforced symmetry: I2. Periphery (PX only) is absent/oversym. Core is structured. Interior is oversymm. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.12 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
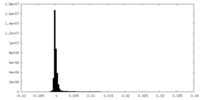
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Half-map 1. Fungal PDC (N. crassa). Recombinant preparation-tE2...
| File | emd_11270_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1. Fungal PDC (N. crassa). Recombinant preparation-tE2 PX30. Enforced symmetry: I2. Periphery (PX only) is absent/oversym. Core is structured. Interior is oversymm. | ||||||||||||
| Projections & Slices |
| ||||||||||||
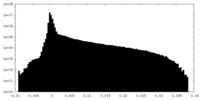
| Density Histograms |
-Half map: Half-map 2. Fungal PDC (N. crassa). Recombinant preparation-tE2...
| File | emd_11270_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2. Fungal PDC (N. crassa). Recombinant preparation-tE2 PX30. Enforced symmetry: I2. Periphery (PX only) is absent/oversym. Core is structured. Interior is oversymm. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Core of Pyruvate dehydrogenase complex with asymmetric interior P...
| Entire | Name: Core of Pyruvate dehydrogenase complex with asymmetric interior PX component. |
|---|---|
| Components |
|
-Supramolecule #1: Core of Pyruvate dehydrogenase complex with asymmetric interior P...
| Supramolecule | Name: Core of Pyruvate dehydrogenase complex with asymmetric interior PX component. type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) |
-Macromolecule #1: Dihydrolipoyllysine-residue acetyltransferase component of pyruva...
| Macromolecule | Name: Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex, mitochondrial type: protein_or_peptide / ID: 1 / Number of copies: 60 / Enantiomer: LEVO / EC number: dihydrolipoyllysine-residue acetyltransferase |
|---|---|
| Source (natural) | Organism:  Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) (fungus) |
| Molecular weight | Theoretical: 30.083316 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRGSHHHHHH GMASMTGGQQ MGRDLYDDDD KDRWGSLVPR GSHMAAYTDV PISGMRKTIA ARLKESVTEN PHFFVSTNLS VSKLLKLRQ ALNSSADGRY KLSVNDFLIK AMGIASKRVP TVNSSWRDGV IRQFETVDVS VAVATPNGLI TPIVKGVEGK G LESISAAV ...String: MRGSHHHHHH GMASMTGGQQ MGRDLYDDDD KDRWGSLVPR GSHMAAYTDV PISGMRKTIA ARLKESVTEN PHFFVSTNLS VSKLLKLRQ ALNSSADGRY KLSVNDFLIK AMGIASKRVP TVNSSWRDGV IRQFETVDVS VAVATPNGLI TPIVKGVEGK G LESISAAV KELAKKARDG KLKPEEYQGG SISISNMGMN PAVQSFTAII NPPQAAILAV GAPQKVAVPV ENEDGTTGVS WD EQIIVTA SFDHKVVDGA VGAEWIRELK KVIENPLELL L UniProtKB: Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex, mitochondrial |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 27.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6zlo: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)