[English] 日本語
 Yorodumi
Yorodumi- EMDB-24625: Archaeal DNA ligase and heterotrimeric PCNA in complex with end-j... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24625 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Archaeal DNA ligase and heterotrimeric PCNA in complex with end-joined DNA | |||||||||
 Map data Map data | Sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA ligase / PCNA / cryo-EM / LIGASE-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationDNA ligase (ATP) / DNA ligase (ATP) activity / lagging strand elongation / DNA biosynthetic process / leading strand elongation / DNA polymerase processivity factor activity / regulation of DNA replication / DNA recombination / cell division / DNA repair ...DNA ligase (ATP) / DNA ligase (ATP) activity / lagging strand elongation / DNA biosynthetic process / leading strand elongation / DNA polymerase processivity factor activity / regulation of DNA replication / DNA recombination / cell division / DNA repair / DNA binding / ATP binding / metal ion binding Similarity search - Function | |||||||||
| Biological species |   Saccharolobus solfataricus (archaea) / Saccharolobus solfataricus (archaea) /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.2 Å | |||||||||
 Authors Authors | Sverzhinsky A / Pascal JM | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2022 Journal: Structure / Year: 2022Title: Cryo-EM structures and biochemical insights into heterotrimeric PCNA regulation of DNA ligase. Authors: Aleksandr Sverzhinsky / Alan E Tomkinson / John M Pascal /   Abstract: DNA ligases act in the final step of many DNA repair pathways and are commonly regulated by the DNA sliding clamp proliferating cell nuclear antigen (PCNA), but there are limited insights into the ...DNA ligases act in the final step of many DNA repair pathways and are commonly regulated by the DNA sliding clamp proliferating cell nuclear antigen (PCNA), but there are limited insights into the physical basis for this regulation. Here, we use single-particle cryoelectron microscopy (cryo-EM) to analyze an archaeal DNA ligase and heterotrimeric PCNA in complex with a single-strand DNA break. The cryo-EM structures highlight a continuous DNA-binding surface formed between DNA ligase and PCNA that supports the distorted conformation of the DNA break undergoing repair and contributes to PCNA stimulation of DNA ligation. DNA ligase is conformationally flexible within the complex, with its domains fully ordered only when encircling the repaired DNA to form a stacked ring structure with PCNA. The structures highlight DNA ligase structural transitions while docked on PCNA, changes in DNA conformation during ligation, and the potential for DNA ligase domains to regulate PCNA accessibility to other repair factors. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24625.map.gz emd_24625.map.gz | 22 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24625-v30.xml emd-24625-v30.xml emd-24625.xml emd-24625.xml | 22.6 KB 22.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_24625.png emd_24625.png | 135 KB | ||
| Filedesc metadata |  emd-24625.cif.gz emd-24625.cif.gz | 7.1 KB | ||
| Others |  emd_24625_additional_1.map.gz emd_24625_additional_1.map.gz | 22 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24625 http://ftp.pdbj.org/pub/emdb/structures/EMD-24625 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24625 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24625 | HTTPS FTP |
-Related structure data
| Related structure data |  7rpxMC  7rpoC  7rpwC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_24625.map.gz / Format: CCP4 / Size: 24.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24625.map.gz / Format: CCP4 / Size: 24.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Unsharpened map
| File | emd_24625_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
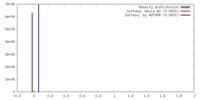
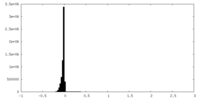
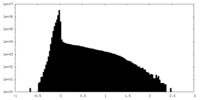
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of DNA Ligase with PCNA1-2-3 and end-joined DNA
| Entire | Name: Ternary complex of DNA Ligase with PCNA1-2-3 and end-joined DNA |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of DNA Ligase with PCNA1-2-3 and end-joined DNA
| Supramolecule | Name: Ternary complex of DNA Ligase with PCNA1-2-3 and end-joined DNA type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#6 |
|---|---|
| Source (natural) | Organism:   Saccharolobus solfataricus (archaea) Saccharolobus solfataricus (archaea) |
| Molecular weight | Theoretical: 175 KDa |
-Macromolecule #1: DNA polymerase sliding clamp 1
| Macromolecule | Name: DNA polymerase sliding clamp 1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharolobus solfataricus (archaea) / Strain: ATCC 35092 / DSM 1617 / JCM 11322 / P2 Saccharolobus solfataricus (archaea) / Strain: ATCC 35092 / DSM 1617 / JCM 11322 / P2 |
| Molecular weight | Theoretical: 28.711961 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAFKIVYPNA KDFFSFINSI TNVTDSIILN FTEDGIFSRH LTEDKVLMAI MRIPKDVLSE YSIDSPTSVK LDVSSVKKIL SKASSKKAT IELTETDSGL KIIIRDEKSG AKSTIYIKAE KGQVEQLTEP KVNLAVNFTT DESVLNVIAA DVTLVGEEMR I STEEDKIK ...String: MAFKIVYPNA KDFFSFINSI TNVTDSIILN FTEDGIFSRH LTEDKVLMAI MRIPKDVLSE YSIDSPTSVK LDVSSVKKIL SKASSKKAT IELTETDSGL KIIIRDEKSG AKSTIYIKAE KGQVEQLTEP KVNLAVNFTT DESVLNVIAA DVTLVGEEMR I STEEDKIK IEAGEEGKRY VAFLMKDKPL KELSIDTSAS SSYSAEMFKD AVKGLRGFSA PTMVSFGENL PMKIDVEAVS GG HMIFWIA PRLLEHHHHH H UniProtKB: DNA polymerase sliding clamp 1 |
-Macromolecule #2: DNA polymerase sliding clamp 2
| Macromolecule | Name: DNA polymerase sliding clamp 2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharolobus solfataricus (archaea) / Strain: ATCC 35092 / DSM 1617 / JCM 11322 / P2 Saccharolobus solfataricus (archaea) / Strain: ATCC 35092 / DSM 1617 / JCM 11322 / P2 |
| Molecular weight | Theoretical: 27.461084 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKAKVIDAVS FSYILRTVGD FLSEANFIVT KEGIRVSGID PSRVVFLDIF LPSSYFEGFE VSQEKEIIGF KLEDVNDILK RVLKDDTLI LSSNESKLTL TFDGEFTRSF ELPLIQVEST QPPSVNLEFP FKAQLLTITF ADIIDELSDL GEVLNIHSKE N KLYFEVIG ...String: MKAKVIDAVS FSYILRTVGD FLSEANFIVT KEGIRVSGID PSRVVFLDIF LPSSYFEGFE VSQEKEIIGF KLEDVNDILK RVLKDDTLI LSSNESKLTL TFDGEFTRSF ELPLIQVEST QPPSVNLEFP FKAQLLTITF ADIIDELSDL GEVLNIHSKE N KLYFEVIG DLSTAKVELS TDNGTLLEAS GADVSSSYGM EYVANTTKMR RASDSMELYF GSQIPLKLRF KLPQEGYGDF YI APRAD UniProtKB: DNA polymerase sliding clamp 2 |
-Macromolecule #3: DNA polymerase sliding clamp 3
| Macromolecule | Name: DNA polymerase sliding clamp 3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharolobus solfataricus (archaea) / Strain: ATCC 35092 / DSM 1617 / JCM 11322 / P2 Saccharolobus solfataricus (archaea) / Strain: ATCC 35092 / DSM 1617 / JCM 11322 / P2 |
| Molecular weight | Theoretical: 28.560268 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKVVYDDVRV LKDIIQALAR LVDEAVLKFK QDSVELVALD RAHISLISVN LPREMFKEYD VNDEFKFGFN TQYLMKILKV AKRKEAIEI ASESPDSVII NIIGSTNREF NVRNLEVSEQ EIPEINLQFD ISATISSDGF KSAISEVSTV TDNVVVEGHE D RILIKAEG ...String: MKVVYDDVRV LKDIIQALAR LVDEAVLKFK QDSVELVALD RAHISLISVN LPREMFKEYD VNDEFKFGFN TQYLMKILKV AKRKEAIEI ASESPDSVII NIIGSTNREF NVRNLEVSEQ EIPEINLQFD ISATISSDGF KSAISEVSTV TDNVVVEGHE D RILIKAEG ESEVEVEFSK DTGGLQDLEF SKESKNSYSA EYLDDVLSLT KLSDYVKISF GNQKPLQLFF NMEGGGKVTY LL APKVLEH HHHHH UniProtKB: DNA polymerase sliding clamp 3 |
-Macromolecule #6: DNA ligase
| Macromolecule | Name: DNA ligase / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA ligase (ATP) |
|---|---|
| Source (natural) | Organism:   Saccharolobus solfataricus (archaea) / Strain: ATCC 35092 / DSM 1617 / JCM 11322 / P2 Saccharolobus solfataricus (archaea) / Strain: ATCC 35092 / DSM 1617 / JCM 11322 / P2 |
| Molecular weight | Theoretical: 69.998375 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGSSHHHHHH SSGLVPRGSH MEFKVIAEYF DKLEKISSRL QLTALLADLL SKSDKTIIDK VVYIIQGKLW PDFLGYPELG IGEKFLIKA ISIATNTDEN SVENLYKTIG DLGEVARRLK SKQQSTGILG FLGTTSKESL TVDEVYSTLS KVALTTGEGS R DLKIRLLA ...String: MGSSHHHHHH SSGLVPRGSH MEFKVIAEYF DKLEKISSRL QLTALLADLL SKSDKTIIDK VVYIIQGKLW PDFLGYPELG IGEKFLIKA ISIATNTDEN SVENLYKTIG DLGEVARRLK SKQQSTGILG FLGTTSKESL TVDEVYSTLS KVALTTGEGS R DLKIRLLA GLLKKADPLE AKFLVRFVEG RLRVGIGDAT VLDAMAIAFG GGQSASEIIE RAYNLRADLG NIAKIIVEKG IE ALKTLKP QVGIPIRPML AERLSNPEEI LKKMGGNAIV DYKYDGERAQ IHKKEDKIFI FSRRLENITS QYPDVVDYVS KYI EGKEFI IEGEIVAIDP ESGEMRPFQE LMHRKRKSDI YEAIKEYPVN VFLFDLMYYE DVDYTTKPLE ARRKLLESIV KPND YVKIA HHIQANNVED LKSFFYRAIS EGGEGVMVKA IGKDAIYQAG ARGWLWIKLK RDYQSEMADT VDLVVVGGFY GKGKR GGKI SSLLMAAYNP KTDSFESVCK VASGFSDEQL DELQKKLMEI KRDVKHPRVN SKMEPDIWVE PVYVAEIIGS EITISP LHT CCQDVVEKDA GLSIRFPRFI RWRDDKSPED ATTTDEILEM YNKQPKKKIE SPAVDESV UniProtKB: DNA ligase |
-Macromolecule #4: End-joined DNA
| Macromolecule | Name: End-joined DNA / type: dna / ID: 4 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.549299 KDa |
| Sequence | String: (DG)(DT)(DA)(DT)(DC)(DC)(DT)(DC)(DG)(DT) (DA)(DG)(DT)(DG)(DC)(DA)(DG)(DA)(DT)(DG) (DC)(DG)(DT)(DC)(DG)(DT)(DC)(DG)(DG) (DA)(DC)(DT)(DG)(DA)(DT)(DT)(DC)(DG)(DG) (DT) (DA)(DG)(DA)(DT)(DC)(DT)(DG) |
-Macromolecule #5: Template strand DNA
| Macromolecule | Name: Template strand DNA / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.403264 KDa |
| Sequence | String: (DC)(DA)(DG)(DA)(DT)(DC)(DT)(DA)(DC)(DC) (DG)(DA)(DA)(DT)(DC)(DA)(DG)(DT)(DC)(DC) (DG)(DA)(DC)(DG)(DA)(DC)(DG)(DC)(DA) (DT)(DC)(DT)(DG)(DC)(DA)(DC)(DT)(DA)(DC) (DG) (DA)(DG)(DG)(DA)(DT)(DA)(DC) |
-Macromolecule #7: MANGANESE (II) ION
| Macromolecule | Name: MANGANESE (II) ION / type: ligand / ID: 7 / Number of copies: 3 / Formula: MN |
|---|---|
| Molecular weight | Theoretical: 54.938 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.35 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 15 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: wait time 0, blot force 1, blot time 1, drain time 0. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 2 / Number real images: 8060 / Average electron dose: 100.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 4.2 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC (ver. 3.1) / Number images used: 192490 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 3.1) |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC (ver. 3.1) |
-Atomic model buiding 1
| Initial model |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Protocol: RIGID BODY FIT | ||||||||||||
| Output model |  PDB-7rpx: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)