+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8474 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human CTP synthase filament | |||||||||
 Map data Map data | Human CTP synthase filament | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | nucleotide metabolism / enzyme / filament / LIGASE / PROTEIN FIBRIL | |||||||||
| Function / homology |  Function and homology information Function and homology informationhistone H1N76/N77 asparagine deamidase activity / protein asparagine deamidase activity / CTP synthase (glutamine hydrolysing) / CTP synthase activity / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / negative regulation of chromosome condensation / Interconversion of nucleotide di- and triphosphates / CTP biosynthetic process ...histone H1N76/N77 asparagine deamidase activity / protein asparagine deamidase activity / CTP synthase (glutamine hydrolysing) / CTP synthase activity / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / negative regulation of chromosome condensation / Interconversion of nucleotide di- and triphosphates / CTP biosynthetic process / nucleobase-containing compound metabolic process / Hydrolases; Acting on carbon-nitrogen bonds, other than peptide bonds; In linear amides / negative regulation of type I interferon production / DNA repair-dependent chromatin remodeling / B cell proliferation / T cell proliferation / response to xenobiotic stimulus / chromatin / ATP binding / membrane / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | helical reconstruction / cryo EM / Resolution: 6.1 Å | |||||||||
 Authors Authors | Lynch EM / Kollman JM | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2017 Journal: Nat Struct Mol Biol / Year: 2017Title: Human CTP synthase filament structure reveals the active enzyme conformation. Authors: Eric M Lynch / Derrick R Hicks / Matthew Shepherd / James A Endrizzi / Allison Maker / Jesse M Hansen / Rachael M Barry / Zemer Gitai / Enoch P Baldwin / Justin M Kollman /   Abstract: The universally conserved enzyme CTP synthase (CTPS) forms filaments in bacteria and eukaryotes. In bacteria, polymerization inhibits CTPS activity and is required for nucleotide homeostasis. Here we ...The universally conserved enzyme CTP synthase (CTPS) forms filaments in bacteria and eukaryotes. In bacteria, polymerization inhibits CTPS activity and is required for nucleotide homeostasis. Here we show that for human CTPS, polymerization increases catalytic activity. The cryo-EM structures of bacterial and human CTPS filaments differ considerably in overall architecture and in the conformation of the CTPS protomer, explaining the divergent consequences of polymerization on activity. The structure of human CTPS filament, the first structure of the full-length human enzyme, reveals a novel active conformation. The filament structures elucidate allosteric mechanisms of assembly and regulation that rely on a conserved conformational equilibrium. The findings may provide a mechanism for increasing human CTPS activity in response to metabolic state and challenge the assumption that metabolic filaments are generally storage forms of inactive enzymes. Allosteric regulation of CTPS polymerization by ligands likely represents a fundamental mechanism underlying assembly of other metabolic filaments. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8474.map.gz emd_8474.map.gz | 201 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8474-v30.xml emd-8474-v30.xml emd-8474.xml emd-8474.xml | 11.1 KB 11.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8474.png emd_8474.png | 109.2 KB | ||
| Filedesc metadata |  emd-8474.cif.gz emd-8474.cif.gz | 5.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8474 http://ftp.pdbj.org/pub/emdb/structures/EMD-8474 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8474 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8474 | HTTPS FTP |
-Related structure data
| Related structure data |  5u03MC  8475C  8476C  8490C  8491C  8504C  8513C  5tkvC  5u05C  5u3cC  5u6rC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8474.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8474.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human CTP synthase filament | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.26 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
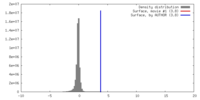
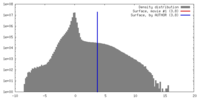
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human CTP synthase 1 filament
| Entire | Name: human CTP synthase 1 filament |
|---|---|
| Components |
|
-Supramolecule #1: human CTP synthase 1 filament
| Supramolecule | Name: human CTP synthase 1 filament / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: hCTPS1 filaments assembled in the presence of UTP, ATP, and GTP |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: CTP synthase 1
| Macromolecule | Name: CTP synthase 1 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: CTP synthase (glutamine hydrolysing) |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 66.777297 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKYILVTGGV ISGIGKGIIA SSVGTILKSC GLHVTSIKID PYINIDAGTF SPYEHGEVFV LDDGGEVDLD LGNYERFLDI RLTKDNNLT TGKIYQYVIN KERKGDYLGK TVQVVPHITD AIQEWVMRQA LIPVDEDGLE PQVCVIELGG TVGDIESMPF I EAFRQFQF ...String: MKYILVTGGV ISGIGKGIIA SSVGTILKSC GLHVTSIKID PYINIDAGTF SPYEHGEVFV LDDGGEVDLD LGNYERFLDI RLTKDNNLT TGKIYQYVIN KERKGDYLGK TVQVVPHITD AIQEWVMRQA LIPVDEDGLE PQVCVIELGG TVGDIESMPF I EAFRQFQF KVKRENFCNI HVSLVPQPSS TGEQKTKPTQ NSVRELRGLG LSPDLVVCRC SNPLDTSVKE KISMFCHVEP EQ VICVHDV SSIYRVPLLL EEQGVVDYFL RRLDLPIERQ PRKMLMKWKE MADRYDRLLE TCSIALVGKY TKFSDSYASV IKA LEHSAL AINHKLEIKY IDSADLEPIT SQEEPVRYHE AWQKLCSAHG VLVPGGFGVR GTEGKIQAIA WARNQKKPFL GVCL GMQLA VVEFSRNVLG WQDANSTEFD PTTSHPVVVD MPEHNPGQMG GTMRLGKRRT LFQTKNSVMR KLYGDADYLE ERHRH RFEV NPVWKKCLEE QGLKFVGQDV EGERMEIVEL EDHPFFVGVQ YHPEFLSRPI KPSPPYFGLL LASVGRLSHY LQKGCR LSP RDTYSDRSGS SSPDSEITEL KFPSINHD UniProtKB: CTP synthase 1 |
-Macromolecule #2: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 2 / Number of copies: 4 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Macromolecule #3: URIDINE 5'-TRIPHOSPHATE
| Macromolecule | Name: URIDINE 5'-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 4 / Formula: UTP |
|---|---|
| Molecular weight | Theoretical: 484.141 Da |
| Chemical component information |  ChemComp-UTP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 7.9 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Δz: 104.1 Å Applied symmetry - Helical parameters - Δ&Phi: 60.6 ° Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Resolution.type: BY AUTHOR / Resolution: 6.1 Å / Resolution method: FSC 0.143 CUT-OFF / Software: (Name: SPIDER, hsearch_lorentz, CTFFIND) / Number images used: 24880 |
|---|---|
| Startup model | Type of model: OTHER / Details: Cylinder |
| Final angle assignment | Type: NOT APPLICABLE |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)