[English] 日本語
 Yorodumi
Yorodumi- PDB-5a0y: METHYL-COENZYME M REDUCTASE FROM METHANOTHERMOBACTER MARBURGENSIS... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5a0y | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
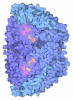
| Title | METHYL-COENZYME M REDUCTASE FROM METHANOTHERMOBACTER MARBURGENSIS AT 1.1 A RESOLUTION | |||||||||
 Components Components | (METHYL-COENZYME M REDUCTASE I SUBUNIT ...) x 3 | |||||||||
 Keywords Keywords | TRANSFERASE / POST-TRANSLATIONAL MODIFICATION / BINDING SITES / CATALYSIS / COENZYMES / DISULFIDES / HYDROGEN / HYDROGEN BONDING / LIGANDS / MESNA / METALLOPORPHYRINS / METHANE / METHANOBACTERIUM / MODELS / MOLECULAR / NICKEL / OXIDATION-REDUCTION / OXIDOREDUCTASES / PHOSPHOTHREONINE / PROTEIN CONFORMATION / PROTEIN FOLDING / PROTEIN STRUCTURE | |||||||||
| Function / homology |  Function and homology information Function and homology informationcoenzyme-B sulfoethylthiotransferase / coenzyme-B sulfoethylthiotransferase activity / methanogenesis / metal ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |   METHANOTHERMOBACTER MARBURGENSIS (archaea) METHANOTHERMOBACTER MARBURGENSIS (archaea) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.1 Å MOLECULAR REPLACEMENT / Resolution: 1.1 Å | |||||||||
 Authors Authors | Wagner, T. / Ermler, U. | |||||||||
 Citation Citation |  Journal: Angew.Chem.Int.Ed.Engl. / Year: 2016 Journal: Angew.Chem.Int.Ed.Engl. / Year: 2016Title: Didehydroaspartate Modification in Methyl-Coenzyme M Reductase Catalyzing Methane Formation. Authors: Wagner, T. / Kahnt, J. / Ermler, U. / Shima, S. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5a0y.cif.gz 5a0y.cif.gz | 1.1 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5a0y.ent.gz pdb5a0y.ent.gz | 904.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5a0y.json.gz 5a0y.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/a0/5a0y https://data.pdbj.org/pub/pdb/validation_reports/a0/5a0y ftp://data.pdbj.org/pub/pdb/validation_reports/a0/5a0y ftp://data.pdbj.org/pub/pdb/validation_reports/a0/5a0y | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5a8kC  5a8rC  5a8wC  3potS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||||||
| Unit cell |
| ||||||||||||||||
| Noncrystallographic symmetry (NCS) | NCS oper:
|
- Components
Components
-METHYL-COENZYME M REDUCTASE I SUBUNIT ... , 3 types, 6 molecules ADBECF
| #1: Protein | Mass: 60637.648 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Details: GERMAN COLLECTION OF MICROORGANISMS (DSM) Source: (natural)   METHANOTHERMOBACTER MARBURGENSIS (archaea) METHANOTHERMOBACTER MARBURGENSIS (archaea)Strain: MARBURG References: UniProt: P11558, coenzyme-B sulfoethylthiotransferase #2: Protein | Mass: 47279.676 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Details: GERMAN COLLECTION OF MICROORGANISMS (DSM) Source: (natural)   METHANOTHERMOBACTER MARBURGENSIS (archaea) METHANOTHERMOBACTER MARBURGENSIS (archaea)Strain: MARBURG References: UniProt: P11560, coenzyme-B sulfoethylthiotransferase #3: Protein | Mass: 28797.234 Da / Num. of mol.: 2 / Source method: isolated from a natural source / Details: GERMAN COLLECTION OF MICROORGANISMS (DSM) Source: (natural)   METHANOTHERMOBACTER MARBURGENSIS (archaea) METHANOTHERMOBACTER MARBURGENSIS (archaea)Strain: MARBURG References: UniProt: P11562, coenzyme-B sulfoethylthiotransferase |
|---|
-Non-polymers , 8 types, 2821 molecules 











| #4: Chemical | ChemComp-MG / #5: Chemical | #6: Chemical | #7: Chemical | #8: Chemical | ChemComp-K / | #9: Chemical | #10: Chemical | #11: Water | ChemComp-HOH / | |
|---|
-Details
| Nonpolymer details | DIDEHYDROASPARTATE (DYA): DIDEHYDROASPARTATE IS AN ASPARTATE CONTAINING A DOUBLE BOND BETWEEN THE C ...DIDEHYDROA |
|---|---|
| Sequence details | RESIDUE 450 IS A DIDEHYDROA |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.18 Å3/Da / Density % sol: 43.65 % / Description: NONE |
|---|---|
| Crystal grow | pH: 9 Details: 27.5% (V/V) PEG 400, 100 MM HEPES PH 7.5, 250 MM MGCL2, AND 200 MM NACL |
-Data collection
| Diffraction | Mean temperature: 77 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SOLEIL SOLEIL  / Beamline: PROXIMA 1 / Wavelength: 0.85506 / Beamline: PROXIMA 1 / Wavelength: 0.85506 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Dec 11, 2014 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.85506 Å / Relative weight: 1 |
| Reflection | Resolution: 1.1→48.35 Å / Num. obs: 937928 / % possible obs: 99.3 % / Observed criterion σ(I): 1.7 / Redundancy: 3.4 % / Biso Wilson estimate: 9.06 Å2 / Rmerge(I) obs: 0.06 / Net I/σ(I): 10.5 |
| Reflection shell | Resolution: 1.1→1.16 Å / Redundancy: 3.3 % / Rmerge(I) obs: 0.7 / Mean I/σ(I) obs: 1.7 / % possible all: 97.8 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 3POT Resolution: 1.1→48.346 Å / SU ML: 0.1 / σ(F): 1.33 / Phase error: 12.26 / Stereochemistry target values: ML Details: CONSIDERING THE RESOLUTION HYDROGEN REFINEMENT MODEL HAS BEEN USED WITH THE RIDING MODEL A NEW POST-TRANSLATIONAL MODIFICATION IS PRESENT IN THE CHAIN A AND D CALLED DIDEHYDROASPARTATE AT THE POSITION 450
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 15.13 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.1→48.346 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj