+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-4057 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of the yeast spliceosome immediately after branching. 3D class containing helicase module. | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | spliceosome / snRNP / pre-mRNA splicing / trans-esterification / lariat intermediate / complex C / splicing | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報RNA exon ligation / snRNA metabolic process / snRNA modification / post-spliceosomal complex / U2-type post-mRNA release spliceosomal complex / cellular bud site selection / post-mRNA release spliceosomal complex / 3'-5' RNA helicase activity / generation of catalytic spliceosome for first transesterification step / cis assembly of pre-catalytic spliceosome ...RNA exon ligation / snRNA metabolic process / snRNA modification / post-spliceosomal complex / U2-type post-mRNA release spliceosomal complex / cellular bud site selection / post-mRNA release spliceosomal complex / 3'-5' RNA helicase activity / generation of catalytic spliceosome for first transesterification step / cis assembly of pre-catalytic spliceosome / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / U4/U6 snRNP / 7-methylguanosine cap hypermethylation / U2-type catalytic step 1 spliceosome / pre-mRNA binding / ATP-dependent activity, acting on RNA / pICln-Sm protein complex / snRNP binding / small nuclear ribonucleoprotein complex / splicing factor binding / SMN-Sm protein complex / spliceosomal tri-snRNP complex / mRNA cis splicing, via spliceosome / commitment complex / U2-type prespliceosome assembly / U2-type spliceosomal complex / U2-type catalytic step 2 spliceosome / U2 snRNP / U1 snRNP / U4 snRNP / U2-type prespliceosome / poly(U) RNA binding / precatalytic spliceosome / generation of catalytic spliceosome for second transesterification step / Formation of TC-NER Pre-Incision Complex / mRNA 5'-splice site recognition / spliceosomal complex assembly / mRNA 3'-splice site recognition / Gap-filling DNA repair synthesis and ligation in TC-NER / spliceosomal tri-snRNP complex assembly / DNA replication origin binding / Prp19 complex / Dual incision in TC-NER / U5 snRNA binding / DNA replication initiation / U5 snRNP / protein K63-linked ubiquitination / U2 snRNA binding / U6 snRNA binding / pre-mRNA intronic binding / spliceosomal snRNP assembly / U1 snRNA binding / U4/U6 x U5 tri-snRNP complex / positive regulation of cell cycle / catalytic step 2 spliceosome / RNA splicing / positive regulation of RNA splicing / spliceosomal complex / mRNA splicing, via spliceosome / RING-type E3 ubiquitin transferase / metallopeptidase activity / ubiquitin-protein transferase activity / ubiquitin protein ligase activity / nucleic acid binding / RNA helicase activity / RNA helicase / DNA repair / GTPase activity / mRNA binding / chromatin binding / chromatin / GTP binding / ATP hydrolysis activity / mitochondrion / DNA binding / RNA binding / zinc ion binding / ATP binding / metal ion binding / identical protein binding / nucleus / cytosol / cytoplasm 類似検索 - 分子機能 | |||||||||
| 生物種 |  | |||||||||
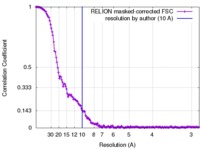
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 10.0 Å | |||||||||
 データ登録者 データ登録者 | Galej WP / Wilkinson MF | |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2016 ジャーナル: Nature / 年: 2016タイトル: Cryo-EM structure of the spliceosome immediately after branching. 著者: Wojciech P Galej / Max E Wilkinson / Sebastian M Fica / Chris Oubridge / Andrew J Newman / Kiyoshi Nagai /  要旨: Precursor mRNA (pre-mRNA) splicing proceeds by two consecutive transesterification reactions via a lariat-intron intermediate. Here we present the 3.8 Å cryo-electron microscopy structure of the ...Precursor mRNA (pre-mRNA) splicing proceeds by two consecutive transesterification reactions via a lariat-intron intermediate. Here we present the 3.8 Å cryo-electron microscopy structure of the spliceosome immediately after lariat formation. The 5'-splice site is cleaved but remains close to the catalytic Mg site in the U2/U6 small nuclear RNA (snRNA) triplex, and the 5'-phosphate of the intron nucleotide G(+1) is linked to the branch adenosine 2'OH. The 5'-exon is held between the Prp8 amino-terminal and linker domains, and base-pairs with U5 snRNA loop 1. Non-Watson-Crick interactions between the branch helix and 5'-splice site dock the branch adenosine into the active site, while intron nucleotides +3 to +6 base-pair with the U6 snRNA ACAGAGA sequence. Isy1 and the step-one factors Yju2 and Cwc25 stabilize docking of the branch helix. The intron downstream of the branch site emerges between the Prp8 reverse transcriptase and linker domains and extends towards the Prp16 helicase, suggesting a plausible mechanism of remodelling before exon ligation. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_4057.map.gz emd_4057.map.gz | 245.2 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-4057-v30.xml emd-4057-v30.xml emd-4057.xml emd-4057.xml | 60 KB 60 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_4057_fsc.xml emd_4057_fsc.xml | 14.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_4057.png emd_4057.png | 47 KB | ||
| Filedesc metadata |  emd-4057.cif.gz emd-4057.cif.gz | 17.6 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4057 http://ftp.pdbj.org/pub/emdb/structures/EMD-4057 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4057 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4057 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  5lj5MC  4055C  4056C  4058C  4059C  5lj3C M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | |
| 電子顕微鏡画像生データ |  EMPIAR-10687 (タイトル: Yeast C, Ci, C*, and P complex spliceosomes / Data size: 8.9 TB EMPIAR-10687 (タイトル: Yeast C, Ci, C*, and P complex spliceosomes / Data size: 8.9 TBData #1: Unaligned movies of C-complex spliceosome with 3' splice site AG to AC mutation (Dataset 1) [micrographs - multiframe] Data #2: Unaligned movies of C and C*-complex spliceosomes with 3' splice site AG to AdG mutation (Dataset 2) [micrographs - multiframe] Data #3: Unaligned movies of C and C*-complex spliceosomes with 3' splice site AG to AdG mutation (Dataset 3) [micrographs - multiframe] Data #4: Aligned movies of C-complex spliceosomes with cold-sensitive prp16-302 mutation, purified with Cwc25 (Dataset 4) [micrographs - multiframe] Data #5: Unaligned movies of C-complex spliceosomes with cold-sensitive prp16-302 mutation, purified with Cwc25 and incubated with ATP and Mg (Dataset 5) [micrographs - multiframe] Data #6: Unaligned movies of C, C*, and P-complex spliceosomes with dominant-negative Prp22 mutation K512A, purified with Slu7 (Dataset 6) [micrographs - multiframe] Data #7: Unaligned movies of P-complex spliceosomes with dominant-negative Prp22 mutation K512A, treated with anti-3'exon RNaseH oligo, purified in presence of Mg (Dataset 9) [micrographs - single frame] Data #8: Selected C-complex particles after polishing [picked particles - single frame - processed] Data #9: Selected P-complex particles after polishing [picked particles - single frame - processed] Data #10: Various signal subtractions for C- and P-complex spliceosomes [picked particles - single frame - processed]) |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_4057.map.gz / 形式: CCP4 / 大きさ: 266.8 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_4057.map.gz / 形式: CCP4 / 大きさ: 266.8 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.43 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : Spliceosome immediately after branching
+超分子 #1: Spliceosome immediately after branching
+分子 #1: U5 snRNA (small nuclear RNA)
+分子 #2: Exon 1 (5' exon) of UBC4 pre-mRNA
+分子 #3: Intron of UBC4 pre-mRNA
+分子 #4: U2 snRNA (small nuclear RNA)
+分子 #5: U6 snRNA (small nuclear RNA)
+分子 #6: Pre-mRNA-splicing factor 8
+分子 #7: Pre-mRNA-splicing helicase BRR2
+分子 #8: Protein CWC16
+分子 #9: Pre-mRNA-splicing factor CWC25
+分子 #10: Pre-mRNA-splicing factor SNU114
+分子 #11: Pre-mRNA-splicing factor ISY1
+分子 #12: CWC22
+分子 #13: Pre-mRNA-splicing factor PRP46
+分子 #14: Pre-mRNA-processing protein 45
+分子 #15: Pre-mRNA-splicing factor BUD31
+分子 #16: Pre-mRNA-splicing factor CWC2
+分子 #17: Pre-mRNA-splicing factor SLT11
+分子 #18: Pre-mRNA-splicing factor CEF1
+分子 #19: CWC15
+分子 #20: Pre-mRNA-splicing factor CWC21
+分子 #21: Pre-mRNA-splicing factor CLF1
+分子 #22: Pre-mRNA-splicing factor SYF1
+分子 #23: Small nuclear ribonucleoprotein-associated protein B
+分子 #24: Small nuclear ribonucleoprotein Sm D3
+分子 #25: Small nuclear ribonucleoprotein E
+分子 #26: Small nuclear ribonucleoprotein F
+分子 #27: Small nuclear ribonucleoprotein G
+分子 #28: Small nuclear ribonucleoprotein Sm D1
+分子 #29: Small nuclear ribonucleoprotein Sm D2
+分子 #30: U2 small nuclear ribonucleoprotein A'
+分子 #31: U2 small nuclear ribonucleoprotein B''
+分子 #32: Pre-mRNA-splicing factor ATP-dependent RNA helicase PRP16
+分子 #33: Pre-mRNA-processing factor 19
+分子 #34: Pre-mRNA-splicing factor SNT309
+分子 #35: unknown
+分子 #36: MAGNESIUM ION
+分子 #37: ZINC ION
+分子 #38: GUANOSINE-5'-TRIPHOSPHATE
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 7.8 構成要素:
| ||||||||||||
| グリッド | モデル: Quantifoil R2/2 / 材質: COPPER / メッシュ: 400 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: CONTINUOUS / 支持フィルム - Film thickness: 6 / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 時間: 30 sec. / 前処理 - 雰囲気: AIR | ||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 277 K / 装置: FEI VITROBOT MARK III 詳細: 3 microlitres sample were applied to the grid, left for 30 seconds and then blotted for 2.5-3.0 seconds before plunging.. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 特殊光学系 | エネルギーフィルター - 名称: GIF Quantum |
| 撮影 | フィルム・検出器のモデル: GATAN K2 QUANTUM (4k x 4k) 検出モード: SUPER-RESOLUTION / デジタル化 - 画像ごとのフレーム数: 1-20 / 実像数: 2213 / 平均露光時間: 0.8 sec. / 平均電子線量: 2.0 e/Å2 詳細: Total dose: 40 electrons/Angstrom^2 over 16 seconds. 20 movie frames collected at 1.25 frames per second. |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 倍率(補正後): 35714 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 4.0 µm / 最小 デフォーカス(公称値): 0.5 µm / 倍率(公称値): 81000 |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 詳細 | Used secondary structure restraints generated in ProSMART and LibG. |
|---|---|
| 精密化 | 空間: RECIPROCAL / プロトコル: OTHER |
| 得られたモデル |  PDB-5lj5: |
 ムービー
ムービー コントローラー
コントローラー






























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)