[English] 日本語
 Yorodumi
Yorodumi- EMDB-6413: Cryo-EM structure of the yeast spliceosome at 3.6 angstrom resolution -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6413 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the yeast spliceosome at 3.6 angstrom resolution | |||||||||
 Map data Map data | Reconstruction of spliceosome | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationnucleolar peripheral inclusion body / mRNA Splicing - Major Pathway / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / spliceosomal conformational changes to generate catalytic conformation / siRNA-mediated pericentric heterochromatin formation / post-mRNA release spliceosomal complex / generation of catalytic spliceosome for first transesterification step / U2-type catalytic step 1 spliceosome ...nucleolar peripheral inclusion body / mRNA Splicing - Major Pathway / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / spliceosomal conformational changes to generate catalytic conformation / siRNA-mediated pericentric heterochromatin formation / post-mRNA release spliceosomal complex / generation of catalytic spliceosome for first transesterification step / U2-type catalytic step 1 spliceosome / pre-mRNA binding / pICln-Sm protein complex / SMN-Sm protein complex / spliceosomal tri-snRNP complex / snRNP binding / commitment complex / mRNA cis splicing, via spliceosome / regulatory ncRNA-mediated gene silencing / U2-type catalytic step 2 spliceosome / U2-type spliceosomal complex / U1 snRNP / U2 snRNP / U4 snRNP / U2-type prespliceosome / precatalytic spliceosome / mRNA 5'-splice site recognition / spliceosomal complex assembly / spliceosomal tri-snRNP complex assembly / Prp19 complex / U5 snRNP / U5 snRNA binding / pre-mRNA intronic binding / spliceosomal snRNP assembly / U6 snRNA binding / U2 snRNA binding / protein K63-linked ubiquitination / U1 snRNA binding / pericentric heterochromatin / U4/U6 x U5 tri-snRNP complex / catalytic step 2 spliceosome / peptidylprolyl isomerase / peptidyl-prolyl cis-trans isomerase activity / spliceosomal complex / mRNA splicing, via spliceosome / RING-type E3 ubiquitin transferase / metallopeptidase activity / ubiquitin-protein transferase activity / ubiquitin protein ligase activity / nuclear envelope / protein folding / molecular adaptor activity / cysteine-type deubiquitinase activity / DNA repair / mRNA binding / GTPase activity / GTP binding / DNA binding / RNA binding / zinc ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Yan C / Hang J / Wan R / Huang M / Wong C / Shi Y | |||||||||
 Citation Citation |  Journal: Science / Year: 2015 Journal: Science / Year: 2015Title: Structure of a yeast spliceosome at 3.6-angstrom resolution. Authors: Chuangye Yan / Jing Hang / Ruixue Wan / Min Huang / Catherine C L Wong / Yigong Shi /  Abstract: Splicing of precursor messenger RNA (pre-mRNA) in yeast is executed by the spliceosome, which consists of five small nuclear ribonucleoproteins (snRNPs), NTC (nineteen complex), NTC-related proteins ...Splicing of precursor messenger RNA (pre-mRNA) in yeast is executed by the spliceosome, which consists of five small nuclear ribonucleoproteins (snRNPs), NTC (nineteen complex), NTC-related proteins (NTR), and a number of associated enzymes and cofactors. Here, we report the three-dimensional structure of a Schizosaccharomyces pombe spliceosome at 3.6-angstrom resolution, revealed by means of single-particle cryogenic electron microscopy. This spliceosome contains U2 and U5 snRNPs, NTC, NTR, U6 small nuclear RNA, and an RNA intron lariat. The atomic model includes 10,574 amino acids from 37 proteins and four RNA molecules, with a combined molecular mass of approximately 1.3 megadaltons. Spp42 (Prp8 in Saccharomyces cerevisiae), the key protein component of the U5 snRNP, forms a central scaffold and anchors the catalytic center. Both the morphology and the placement of protein components appear to have evolved to facilitate the dynamic process of pre-mRNA splicing. Our near-atomic-resolution structure of a central spliceosome provides a molecular framework for mechanistic understanding of pre-mRNA splicing. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6413.map.gz emd_6413.map.gz | 166.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6413-v30.xml emd-6413-v30.xml emd-6413.xml emd-6413.xml | 9.4 KB 9.4 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6413.gif 400_6413.gif 80_6413.gif 80_6413.gif | 59.3 KB 4.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6413 http://ftp.pdbj.org/pub/emdb/structures/EMD-6413 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6413 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6413 | HTTPS FTP |
-Related structure data
| Related structure data |  3jb9MC  6414C  6415C  6416C  6417C  6418C  6419C  6420C  6421C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6413.map.gz / Format: CCP4 / Size: 173.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6413.map.gz / Format: CCP4 / Size: 173.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of spliceosome | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
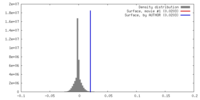
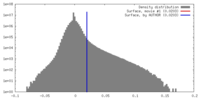
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : spliceosome
| Entire | Name: spliceosome |
|---|---|
| Components |
|
-Supramolecule #1000: spliceosome
| Supramolecule | Name: spliceosome / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Macromolecule #1: Spliceosome
| Macromolecule | Name: Spliceosome / type: protein_or_peptide / ID: 1 / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 75mM NaCl, 10mM Tris-HCL,1mM Mg(OAc)2,1mM imidazole, 0.01% NP40, 1mM TCEP, 2mM EGTA |
| Grid | Details: carbon coated grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV / Method: Blot for 2.5 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Date | Mar 29, 2015 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 (4k x 4k) / Number real images: 2246 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 1.4 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.6 Å / Resolution method: OTHER / Software - Name: RELION / Number images used: 112795 |
|---|
 Movie
Movie Controller
Controller






















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)