[English] 日本語
 Yorodumi
Yorodumi- EMDB-10861: Cryo-EM structure of Tetrahymena thermophila mitochondrial ATP sy... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10861 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Tetrahymena thermophila mitochondrial ATP synthase - F1Fo tetramer | ||||||||||||
 Map data Map data | Local-resolution filtered full map of T. thermophila ATP synthase tetramer, Fo-subcomplexes masked | ||||||||||||
 Sample Sample |
| ||||||||||||
| Function / homology |  Function and homology information Function and homology information: / sulfide oxidation, using sulfide:quinone oxidoreductase / sulfide:quinone oxidoreductase activity / cellular lipid metabolic process / carboxylic ester hydrolase activity / : / : / : / : / photosynthetic electron transport in photosystem I ...: / sulfide oxidation, using sulfide:quinone oxidoreductase / sulfide:quinone oxidoreductase activity / cellular lipid metabolic process / carboxylic ester hydrolase activity / : / : / : / : / photosynthetic electron transport in photosystem I / proton-transporting ATP synthase complex, coupling factor F(o) / : / proton transmembrane transporter activity / proton motive force-driven ATP synthesis / chloroplast thylakoid membrane / photosynthetic electron transport in photosystem II / proton-transporting ATP synthase complex, catalytic core F(1) / H+-transporting two-sector ATPase / proton-transporting ATPase activity, rotational mechanism / FAD binding / proton-transporting ATP synthase activity, rotational mechanism / proton motive force-driven mitochondrial ATP synthesis / ADP binding / hydrolase activity / lipid binding / ATP hydrolysis activity / mitochondrion / ATP binding / membrane / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
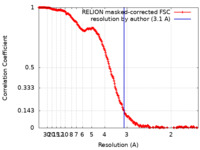
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | ||||||||||||
 Authors Authors | Kock Flygaard R / Muhleip A / Amunts A | ||||||||||||
| Funding support |  Sweden, 3 items Sweden, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Type III ATP synthase is a symmetry-deviated dimer that induces membrane curvature through tetramerization. Authors: Rasmus Kock Flygaard / Alexander Mühleip / Victor Tobiasson / Alexey Amunts /  Abstract: Mitochondrial ATP synthases form functional homodimers to induce cristae curvature that is a universal property of mitochondria. To expand on the understanding of this fundamental phenomenon, we ...Mitochondrial ATP synthases form functional homodimers to induce cristae curvature that is a universal property of mitochondria. To expand on the understanding of this fundamental phenomenon, we characterized the unique type III mitochondrial ATP synthase in its dimeric and tetrameric form. The cryo-EM structure of a ciliate ATP synthase dimer reveals an unusual U-shaped assembly of 81 proteins, including a substoichiometrically bound ATPTT2, 40 lipids, and co-factors NAD and CoQ. A single copy of subunit ATPTT2 functions as a membrane anchor for the dimeric inhibitor IF. Type III specific linker proteins stably tie the ATP synthase monomers in parallel to each other. The intricate dimer architecture is scaffolded by an extended subunit-a that provides a template for both intra- and inter-dimer interactions. The latter results in the formation of tetramer assemblies, the membrane part of which we determined to 3.1 Å resolution. The structure of the type III ATP synthase tetramer and its associated lipids suggests that it is the intact unit propagating the membrane curvature. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10861.map.gz emd_10861.map.gz | 468.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10861-v30.xml emd-10861-v30.xml emd-10861.xml emd-10861.xml | 45.6 KB 45.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10861_fsc.xml emd_10861_fsc.xml | 21.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_10861.png emd_10861.png | 62.4 KB | ||
| Masks |  emd_10861_msk_1.map emd_10861_msk_1.map | 824 MB |  Mask map Mask map | |
| Others |  emd_10861_half_map_1.map.gz emd_10861_half_map_1.map.gz emd_10861_half_map_2.map.gz emd_10861_half_map_2.map.gz | 665.7 MB 665.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10861 http://ftp.pdbj.org/pub/emdb/structures/EMD-10861 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10861 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10861 | HTTPS FTP |
-Validation report
| Summary document |  emd_10861_validation.pdf.gz emd_10861_validation.pdf.gz | 460.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10861_full_validation.pdf.gz emd_10861_full_validation.pdf.gz | 459.4 KB | Display | |
| Data in XML |  emd_10861_validation.xml.gz emd_10861_validation.xml.gz | 24.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10861 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10861 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10861 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10861 | HTTPS FTP |
-Related structure data
| Related structure data |  6ynzMC  6ynvC  6ynwC  6ynxC  6ynyC  6yo0C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10861.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10861.map.gz / Format: CCP4 / Size: 824 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Local-resolution filtered full map of T. thermophila ATP synthase tetramer, Fo-subcomplexes masked | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10861_msk_1.map emd_10861_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map 1
| File | emd_10861_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half-map 2
| File | emd_10861_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half-map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Mitochondrial ATP synthase, F1Fo tetramer
+Supramolecule #1: Mitochondrial ATP synthase, F1Fo tetramer
+Macromolecule #1: Ymf66
+Macromolecule #2: subunit b
+Macromolecule #3: subunit d
+Macromolecule #4: subunit f
+Macromolecule #5: subunit i/j
+Macromolecule #6: subunit k
+Macromolecule #7: Ymf56
+Macromolecule #8: ATPTT3
+Macromolecule #9: ATPTT4
+Macromolecule #10: ATPTT5
+Macromolecule #11: ATPTT6
+Macromolecule #12: ATPTT7
+Macromolecule #13: ATPTT8
+Macromolecule #14: ATPTT9
+Macromolecule #15: ATPTT10
+Macromolecule #16: ATPTT11
+Macromolecule #17: ATPTT12
+Macromolecule #18: ATPTT13
+Macromolecule #19: ATPTT1
+Macromolecule #20: Inhibitor of F1 (IF1)
+Macromolecule #21: ATPTT2
+Macromolecule #22: Oligomycin sensitivity-conferring protein (OSCP)
+Macromolecule #23: ATP synthase subunit gamma
+Macromolecule #24: ATP synthase subunit alpha
+Macromolecule #25: ATP synthase subunit beta
+Macromolecule #26: ATP synthase F0 subunit 9
+Macromolecule #27: subunit delta
+Macromolecule #28: subunit epsilon
+Macromolecule #29: CARDIOLIPIN
+Macromolecule #30: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
+Macromolecule #31: PHOSPHATE ION
+Macromolecule #32: Ubiquinone-8
+Macromolecule #33: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #34: MAGNESIUM ION
+Macromolecule #35: 1,2-Dioleoyl-sn-glycero-3-phosphoethanolamine
+Macromolecule #36: NICOTINAMIDE-ADENINE-DINUCLEOTIDE
+Macromolecule #37: ADENOSINE-5'-DIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.75 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Average electron dose: 30.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal magnification: 165000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
























 Z
Z Y
Y X
X