+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4238 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
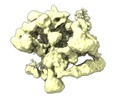
| Title | Human Bact spliceosome state 6 unmasked | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 8.1 Å | |||||||||
 Authors Authors | Haselbach D / Komarov I / Agafonov D / Kastner B / Luehrmann R / Stark H | |||||||||
 Citation Citation |  Journal: Cell / Year: 2018 Journal: Cell / Year: 2018Title: Structure and Conformational Dynamics of the Human Spliceosomal B Complex. Authors: David Haselbach / Ilya Komarov / Dmitry E Agafonov / Klaus Hartmuth / Benjamin Graf / Olexandr Dybkov / Henning Urlaub / Berthold Kastner / Reinhard Lührmann / Holger Stark /  Abstract: The spliceosome is a highly dynamic macromolecular complex that precisely excises introns from pre-mRNA. Here we report the cryo-EM 3D structure of the human B spliceosome at 3.4 Å resolution. In ...The spliceosome is a highly dynamic macromolecular complex that precisely excises introns from pre-mRNA. Here we report the cryo-EM 3D structure of the human B spliceosome at 3.4 Å resolution. In the B state, the spliceosome is activated but not catalytically primed, so that it is functionally blocked prior to the first catalytic step of splicing. The spliceosomal core is similar to the yeast B spliceosome; important differences include the presence of the RNA helicase aquarius and peptidyl prolyl isomerases. To examine the overall dynamic behavior of the purified spliceosome, we developed a principal component analysis-based approach. Calculating the energy landscape revealed eight major conformational states, which we refined to higher resolution. Conformational differences of the highly flexible structural components between these eight states reveal how spliceosomal components contribute to the assembly of the spliceosome, allowing it to generate a dynamic interaction network required for its subsequent catalytic activation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4238.map.gz emd_4238.map.gz | 244.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4238-v30.xml emd-4238-v30.xml emd-4238.xml emd-4238.xml | 13.2 KB 13.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4238.png emd_4238.png | 138.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4238 http://ftp.pdbj.org/pub/emdb/structures/EMD-4238 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4238 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4238 | HTTPS FTP |
-Related structure data
| Related structure data |  4233C  4234C  4235C  4236C  4237C  4239C  4240C  4247C  4248C  4249C  4250C  4251C  4252C  4253C  4254C  4255C  6ff4C  6ff7C C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10160 (Title: Conformational Dynamics of human Bact spliceosome / Data size: 2.8 TB EMPIAR-10160 (Title: Conformational Dynamics of human Bact spliceosome / Data size: 2.8 TBData #1: aligned and summed micrograph stack of human Bact spliceosome [micrographs - single frame] Data #2: aligned, dose-weighted and summed micrograph stack of human Bact spliceosome [micrographs - single frame]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_4238.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4238.map.gz / Format: CCP4 / Size: 282.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.16 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : human Bact spliceosome state 1 unmasked
| Entire | Name: human Bact spliceosome state 1 unmasked |
|---|---|
| Components |
|
-Supramolecule #1: human Bact spliceosome state 1 unmasked
| Supramolecule | Name: human Bact spliceosome state 1 unmasked / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / Strain: HELA Homo sapiens (human) / Strain: HELA |
| Molecular weight | Theoretical: 4.5 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.05 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.9 Component:
| ||||||||||||
| Grid | Model: Quantifoil R3.5/1 / Material: COPPER / Mesh: 400 / Support film - #0 - Film type ID: 1 / Support film - #0 - Material: CARBON / Support film - #0 - topology: HOLEY / Support film - #1 - Film type ID: 2 / Support film - #1 - Material: CARBON / Support film - #1 - topology: CONTINUOUS | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 75 % / Chamber temperature: 277 K / Instrument: LEICA EM GP / Details: blot with blotting sensor. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Alignment procedure | Coma free - Residual tilt: 14.0 mrad |
| Specialist optics | Spherical aberration corrector: Microscope was modified with a Cs corrector with two hexapoles elements |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: INTEGRATING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Sampling interval: 14.0 µm / Number grids imaged: 1 / Number real images: 32000 / Average exposure time: 1.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 0.001 mm / Nominal defocus max: 4.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: RIGID BODY FIT |
|---|
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















