[English] 日本語
 Yorodumi
Yorodumi- EMDB-10522: Cryo-EM structure of Toxoplasma gondii mitochondrial ATP synthase... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10522 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Toxoplasma gondii mitochondrial ATP synthase dimer, peripheral stalk map | |||||||||||||||
 Map data Map data | Toxoplasma gondii ATP synthase dimer, peripheral stalk full map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | mitochondrial / ATP synthase / peripheral stalk / OSCP / MEMBRANE PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationproton transmembrane transporter activity / proton motive force-driven ATP synthesis / proton-transporting ATP synthase complex / proton-transporting ATP synthase activity, rotational mechanism / ADP binding / hydrolase activity / ATP binding / membrane Similarity search - Function | |||||||||||||||
| Biological species |   | |||||||||||||||
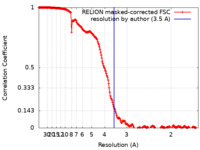
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||||||||
 Authors Authors | Muhleip A / Kock Flygaard R / Amunts A | |||||||||||||||
| Funding support |  Sweden, 4 items Sweden, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: ATP synthase hexamer assemblies shape cristae of Toxoplasma mitochondria. Authors: Alexander Mühleip / Rasmus Kock Flygaard / Jana Ovciarikova / Alice Lacombe / Paula Fernandes / Lilach Sheiner / Alexey Amunts /   Abstract: Mitochondrial ATP synthase plays a key role in inducing membrane curvature to establish cristae. In Apicomplexa causing diseases such as malaria and toxoplasmosis, an unusual cristae morphology has ...Mitochondrial ATP synthase plays a key role in inducing membrane curvature to establish cristae. In Apicomplexa causing diseases such as malaria and toxoplasmosis, an unusual cristae morphology has been observed, but its structural basis is unknown. Here, we report that the apicomplexan ATP synthase assembles into cyclic hexamers, essential to shape their distinct cristae. Cryo-EM was used to determine the structure of the hexamer, which is held together by interactions between parasite-specific subunits in the lumenal region. Overall, we identified 17 apicomplexan-specific subunits, and a minimal and nuclear-encoded subunit-a. The hexamer consists of three dimers with an extensive dimer interface that includes bound cardiolipins and the inhibitor IF. Cryo-ET and subtomogram averaging revealed that hexamers arrange into ~20-megadalton pentagonal pyramids in the curved apical membrane regions. Knockout of the linker protein ATPTG11 resulted in the loss of pentagonal pyramids with concomitant aberrantly shaped cristae. Together, this demonstrates that the unique macromolecular arrangement is critical for the maintenance of cristae morphology in Apicomplexa. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10522.map.gz emd_10522.map.gz | 378.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10522-v30.xml emd-10522-v30.xml emd-10522.xml emd-10522.xml | 21.4 KB 21.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10522_fsc.xml emd_10522_fsc.xml | 19.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_10522.png emd_10522.png | 36 KB | ||
| Masks |  emd_10522_msk_1.map emd_10522_msk_1.map | 669.9 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-10522.cif.gz emd-10522.cif.gz | 6.9 KB | ||
| Others |  emd_10522_half_map_1.map.gz emd_10522_half_map_1.map.gz emd_10522_half_map_2.map.gz emd_10522_half_map_2.map.gz | 538.9 MB 538.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10522 http://ftp.pdbj.org/pub/emdb/structures/EMD-10522 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10522 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10522 | HTTPS FTP |
-Related structure data
| Related structure data |  6tmiMC  6tmgC  6tmhC  6tmjC  6tmkC  6tmlC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10522.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10522.map.gz / Format: CCP4 / Size: 669.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Toxoplasma gondii ATP synthase dimer, peripheral stalk full map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10522_msk_1.map emd_10522_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap 1
| File | emd_10522_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap 2
| File | emd_10522_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Mitochondrial ATP synthase dimer, peripheral stalk
| Entire | Name: Mitochondrial ATP synthase dimer, peripheral stalk |
|---|---|
| Components |
|
-Supramolecule #1: Mitochondrial ATP synthase dimer, peripheral stalk
| Supramolecule | Name: Mitochondrial ATP synthase dimer, peripheral stalk / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 68 KDa |
-Macromolecule #1: ATP synthase subunit alpha
| Macromolecule | Name: ATP synthase subunit alpha / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 61.189168 KDa |
| Sequence | String: MTIHSCLARR AVSVAASGAR AFASGLGARA VAVGALQSAR LLHTSSLRAA GAKISPSEMS RLLEERIAGW KTQTSTEEVG RVVSVGDGI ARLFGLEGVQ AGELVEFQNG MTGMALNLET DNVGVVIFGD DRSVLEGDSV KRTGRIVDVP IGPGLLGRVV D ALGNPIDG ...String: MTIHSCLARR AVSVAASGAR AFASGLGARA VAVGALQSAR LLHTSSLRAA GAKISPSEMS RLLEERIAGW KTQTSTEEVG RVVSVGDGI ARLFGLEGVQ AGELVEFQNG MTGMALNLET DNVGVVIFGD DRSVLEGDSV KRTGRIVDVP IGPGLLGRVV D ALGNPIDG KGPIPAKERR RVELKAPGII PRKSVHEPMM TGLKCVDALV PVGRGQRELI IGDRQTGKTA VAVDAIINQK EI NDSTDDE SKKLYCIYVA VGQKRSTVAQ IVKALEQRDA MKYTTVVAAT ASEAAPLQFL APYSGCAMGE WFRDSGRHCV IIY DDLSKQ ATAYRQMSLL LRRPPGREAY PGDVFYLHSR LLERAAKMGD KSGGGSLTAL PVIETQAGDV SAYIPTNVIS ITDG QIFLE TELFYKGIRP AINVGLSVSR VGSAAQVKAM KQVAGTMKLE LAQYREVAAF AQFGSDLDAS TRQLLTRGTA LTELL KQRQ YSPMKNSVQV CVLYCGVKGY LDPLDPKEIS RFESLFIDYI NANHQDILKT IETEKELSEK TEAKLRAAVD EFVAMN EFK KK UniProtKB: ATP synthase subunit alpha |
-Macromolecule #2: Oligomycin sensitivity conferring protein (OSCP)
| Macromolecule | Name: Oligomycin sensitivity conferring protein (OSCP) / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.669994 KDa |
| Sequence | String: MALPLLASRR LFSSFVFRGQ PSTLSSNLSL VRIRGLHGGS LSPPSATLPR AVQLFSSRIA FSTAAAEDSG ASQTLEGRYA SALFRVAKK KNQLEKVYGD LESVRNALKD SSEFRLFVDS PAVSVQQKLD VLRQLVNRYK FDPLTGNLLT TLVENKRLPM L ARVADAFD ...String: MALPLLASRR LFSSFVFRGQ PSTLSSNLSL VRIRGLHGGS LSPPSATLPR AVQLFSSRIA FSTAAAEDSG ASQTLEGRYA SALFRVAKK KNQLEKVYGD LESVRNALKD SSEFRLFVDS PAVSVQQKLD VLRQLVNRYK FDPLTGNLLT TLVENKRLPM L ARVADAFD AMYRKEKGEV KCLVTSAKPL SAQQQKEIVA ALQNRAGTQA RLIIDYAVSP QIMGGLVVRL GEQVLDFSVA TR LDRLQSQ LLAPL UniProtKB: ATP synthase F1, delta subunit protein |
-Macromolecule #3: subunit d
| Macromolecule | Name: subunit d / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 61.36225 KDa |
| Sequence | String: MQALRRGAAI PSRLLPRRDS WMSLAPFVAP NNAAAWRKLR DGAQEVQTVI ERQSTPGKPQ QIDWAKWESQ IAHKDILNCL KTFYTNQVQ ILDRALGALE TAKTPAPCEG AEKGWALFDA ALSACAKSVE KSEELLSNGA RALWVSCSNP PVWKVNTNEW L DSDQYWQA ...String: MQALRRGAAI PSRLLPRRDS WMSLAPFVAP NNAAAWRKLR DGAQEVQTVI ERQSTPGKPQ QIDWAKWESQ IAHKDILNCL KTFYTNQVQ ILDRALGALE TAKTPAPCEG AEKGWALFDA ALSACAKSVE KSEELLSNGA RALWVSCSNP PVWKVNTNEW L DSDQYWQA FVEKHHFYSQ YQPGVVDPEA PQEVEAFKQA WHSRMGKFND RSDTPMLYAY MNELPSWEYY DLHRSAFLEH MT YFLVRTG GDFRFFPEMP PWQWLAHMEN LRFKLLSVAQ SRRSQLQLAN LERERALDFL PVDVEHHGEE YTQKFLQYET ELF QACAAR LMGHFMFLCD PFIPVQSAEA LSAVTRVDNG KGKLFSLGDD VNALFYLPEQ QRRDVERPTQ AVQTLLGHLE ATGR PFNPC YSELLHVHAE VLEERGEHWL TAPGECVSQA FLRRLRTDDP AYEVYCSYFK EMYERFAGAK EVSMEDGRKR LATIE KNAQ EEAAAYGLAL KTMGSAELAH KAREGAAKLE QLRKAQEKAA GKSAQTVQEN KM UniProtKB: Uncharacterized protein |
-Macromolecule #4: ATPTG12
| Macromolecule | Name: ATPTG12 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.400639 KDa |
| Sequence | String: MLNFIPKRCP SVSLLFGKRP VQRIEVGQAR HQLEIPVETI EKIYEGVDSR LEYHNKDYNA MKWKDFMKLK LDAYHLLEAS QSETAAKSA LSDLNWFSDL ADIYSGQQTM AEMDVALKAQ GEQKLSYPIQ GKNIK UniProtKB: Uncharacterized protein |
-Macromolecule #5: subunit b
| Macromolecule | Name: subunit b / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 64.811602 KDa |
| Sequence | String: MNFSSSARWL AVRQSQTLGH TTRATVAAGR RVLAHSPAAT EFTSFQSLHI GGDVCKLPLA VALGAAPSAL GYGSAKHNQQ RQYATLGSG WSFSKVQYTK YRITKPWTTD TTFDDIILSQ PSKEDFAKFT KEAPLFLRFL KLVTDVEGRQ EAFIQFAKRC E NGLTVEKD ...String: MNFSSSARWL AVRQSQTLGH TTRATVAAGR RVLAHSPAAT EFTSFQSLHI GGDVCKLPLA VALGAAPSAL GYGSAKHNQQ RQYATLGSG WSFSKVQYTK YRITKPWTTD TTFDDIILSQ PSKEDFAKFT KEAPLFLRFL KLVTDVEGRQ EAFIQFAKRC E NGLTVEKD VYVTKKELVD CLWKNGYTDT EINAFEIAFP ADYKFHYPEL AVLFDLTEED CYKYCIRQRA ATPEELVELK YT KPKNLVS SYGLCFLGVW FGLSNTVLSN AWFYSKTFPF GAVFYMLGSY FYRDIREKLW KEEKSLIHTA QENKNMGEES VYK QMKKYA TDTKCLDYLS TFRTEVEDQI ANYKVALVSQ MRRQLTERLV EKLNGIQQAE KLIQGSLQDV MIREIVSSFK DLYK SRPEL HDAAMQSAIQ GLSGSDGAMD PVGAHFKASL QELAKVNLST ATADPMGTVV QRVAAVFQKR EKEFLDTFTV KATEA QEIK TIVDKCHKGN TFDFHALSDE ELRRLEQLYS TVNNRVGFET IHENSIKPVA PLSENSKGFV EFVNTQLEIT KAKLRN ARL TAFAHAFV UniProtKB: Uncharacterized protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL |
|---|---|
| Buffer | pH: 7.5 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV / Details: 3 seconds blot.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Number real images: 4860 / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL |
|---|---|
| Output model |  PDB-6tmi: |
 Movie
Movie Controller
Controller



















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)