+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10191 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of protomer 2 of the ESX-3 core complex | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Type VII Secretion System ESX-3 secretion system T7SS ESX-3 Mycobacterium smegmatis / MEMBRANE PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationHydrolases; Acting on acid anhydrides / hydrolase activity / ATP binding / plasma membrane Similarity search - Function | ||||||||||||||||||
| Biological species |  Mycolicibacterium smegmatis MC2 155 (bacteria) / Mycolicibacterium smegmatis MC2 155 (bacteria) /  Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) | ||||||||||||||||||
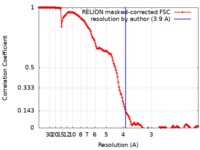
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | ||||||||||||||||||
 Authors Authors | Rivera-Calzada A / Famelis N / Geibel S / Llorca O | ||||||||||||||||||
| Funding support |  Spain, Spain,  Germany, 5 items Germany, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: Architecture of the mycobacterial type VII secretion system. Authors: Nikolaos Famelis / Angel Rivera-Calzada / Gianluca Degliesposti / Maria Wingender / Nicole Mietrach / J Mark Skehel / Rafael Fernandez-Leiro / Bettina Böttcher / Andreas Schlosser / Oscar ...Authors: Nikolaos Famelis / Angel Rivera-Calzada / Gianluca Degliesposti / Maria Wingender / Nicole Mietrach / J Mark Skehel / Rafael Fernandez-Leiro / Bettina Böttcher / Andreas Schlosser / Oscar Llorca / Sebastian Geibel /    Abstract: Host infection by pathogenic mycobacteria, such as Mycobacterium tuberculosis, is facilitated by virulence factors that are secreted by type VII secretion systems. A molecular understanding of the ...Host infection by pathogenic mycobacteria, such as Mycobacterium tuberculosis, is facilitated by virulence factors that are secreted by type VII secretion systems. A molecular understanding of the type VII secretion mechanism has been hampered owing to a lack of three-dimensional structures of the fully assembled secretion apparatus. Here we report the cryo-electron microscopy structure of a membrane-embedded core complex of the ESX-3/type VII secretion system from Mycobacterium smegmatis. The core of the ESX-3 secretion machine consists of four protein components-EccB3, EccC3, EccD3 and EccE3, in a 1:1:2:1 stoichiometry-which form two identical protomers. The EccC3 coupling protein comprises a flexible array of four ATPase domains, which are linked to the membrane through a stalk domain. The domain of unknown function (DUF) adjacent to the stalk is identified as an ATPase domain that is essential for secretion. EccB3 is predominantly periplasmatic, but a small segment crosses the membrane and contacts the stalk domain. This suggests that conformational changes in the stalk domain-triggered by substrate binding at the distal end of EccC3 and subsequent ATP hydrolysis in the DUF-could be coupled to substrate secretion to the periplasm. Our results reveal that the architecture of type VII secretion systems differs markedly from that of other known secretion machines, and provide a structural understanding of these systems that will be useful for the design of antimicrobial strategies that target bacterial virulence. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10191.map.gz emd_10191.map.gz | 255.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10191-v30.xml emd-10191-v30.xml emd-10191.xml emd-10191.xml | 19.9 KB 19.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10191_fsc.xml emd_10191_fsc.xml | 14.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_10191.png emd_10191.png | 65 KB | ||
| Filedesc metadata |  emd-10191.cif.gz emd-10191.cif.gz | 6.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10191 http://ftp.pdbj.org/pub/emdb/structures/EMD-10191 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10191 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10191 | HTTPS FTP |
-Validation report
| Summary document |  emd_10191_validation.pdf.gz emd_10191_validation.pdf.gz | 313.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10191_full_validation.pdf.gz emd_10191_full_validation.pdf.gz | 312.6 KB | Display | |
| Data in XML |  emd_10191_validation.xml.gz emd_10191_validation.xml.gz | 14.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10191 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10191 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10191 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10191 | HTTPS FTP |
-Related structure data
| Related structure data |  6sgzMC  6sgwC  6sgxC  6sgyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10191.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10191.map.gz / Format: CCP4 / Size: 274.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.0635 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
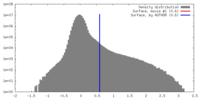
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Protomer2 of the ESX-3 core complex
| Entire | Name: Protomer2 of the ESX-3 core complex |
|---|---|
| Components |
|
-Supramolecule #1: Protomer2 of the ESX-3 core complex
| Supramolecule | Name: Protomer2 of the ESX-3 core complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: ESX-3 core complex The sample consists of four protein components, EccB3:EccC3:EccD3:EccE3 in a 1:1:2:1 stoichiometry Molecular weight of the complex without the amphipol micelle: 0.65 MDa |
|---|---|
| Molecular weight | Theoretical: 650 KDa |
-Supramolecule #2: ESX-3 secretion system protein EccD3
| Supramolecule | Name: ESX-3 secretion system protein EccD3 / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
-Supramolecule #3: ESX-3 secretion system protein EccB3
| Supramolecule | Name: ESX-3 secretion system protein EccB3 / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
-Supramolecule #4: ESX-3 secretion system protein EccC3
| Supramolecule | Name: ESX-3 secretion system protein EccC3 / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
-Supramolecule #5: ESX-3 secretion system protein EccD3
| Supramolecule | Name: ESX-3 secretion system protein EccD3 / type: complex / ID: 5 / Parent: 1 / Macromolecule list: #4 |
|---|---|
| Source (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
-Macromolecule #1: ESX-3 secretion system protein EccD3
| Macromolecule | Name: ESX-3 secretion system protein EccD3 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria)Strain: ATCC 700084 / mc(2)155 |
| Molecular weight | Theoretical: 47.344453 KDa |
| Recombinant expression | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Sequence | String: VMPIVRVAVL AAGDDGGRLT EMALPSELPL REILPAVQRI VQPARENDGA ADPAAAPNPV RLSLAPIGGA PFSLDATLDT VGVVDGDLL ALQAVPSGPP APRIVEDIAD AAVIFSEARR RQWGPTHIAR GAALALIGLI LVGTGLSVAH RVITGDLLGQ F IVSGIALA ...String: VMPIVRVAVL AAGDDGGRLT EMALPSELPL REILPAVQRI VQPARENDGA ADPAAAPNPV RLSLAPIGGA PFSLDATLDT VGVVDGDLL ALQAVPSGPP APRIVEDIAD AAVIFSEARR RQWGPTHIAR GAALALIGLI LVGTGLSVAH RVITGDLLGQ F IVSGIALA TVIAALAVRN RSAVLATSLA VTALVPVAAA FALGVPGDFG APNVLLAAAG VAAWSLISMA GSPDDRGIAV FT ATAVTGV GVLLVAGAAS LWVISSDVIG CALVLLGLIV TVQAAQLSAM WARFPLPVIP APGDPTPAAR PLSVLADLPR RVR VSQAHQ TGVIAAGVLL GVAGSVALVS SANASPWAWY IVVAAAAGAA LRARVWDSAA CKAWLLGHSY LLAVALLVAF VIGD RYQAA LWALAALAVL VLVWIVAALN PKIASPDTYS LPMRRMVGFL ATGLDASLIP VMALLVGLFS LV UniProtKB: ESX-3 secretion system protein EccD3 |
-Macromolecule #2: ESX-3 secretion system ATPase EccB3
| Macromolecule | Name: ESX-3 secretion system ATPase EccB3 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: Hydrolases; Acting on acid anhydrides |
|---|---|
| Source (natural) | Organism:  Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria)Strain: ATCC 700084 / mc(2)155 |
| Molecular weight | Theoretical: 6.579834 KDa |
| Recombinant expression | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Sequence | String: VTRHQVSGWR FVMRRIASGV ALHDTRMLVD PLRTQSRAVL TGALILVTGL VGCFIFSLF UniProtKB: ESX-3 secretion system ATPase EccB3 |
-Macromolecule #3: ESX-3 secretion system protein EccC3
| Macromolecule | Name: ESX-3 secretion system protein EccC3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) |
| Molecular weight | Theoretical: 38.29718 KDa |
| Recombinant expression | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Sequence | String: MSRLIFEHQR RLTPPTTRKG TITIEPPPQL PMRTEEVDAE RADYLRYLSV VRDNVRAHAA EQRAALEWSH PEPEVLATIP GTRRQWERD PRDRDFLVLR AGRHDVPLDA ALKVKDTADE IDLEPVAHSA LRGLLDVQRT VRDAPTGLDV AKLARITVIG E ADEARAAI ...String: MSRLIFEHQR RLTPPTTRKG TITIEPPPQL PMRTEEVDAE RADYLRYLSV VRDNVRAHAA EQRAALEWSH PEPEVLATIP GTRRQWERD PRDRDFLVLR AGRHDVPLDA ALKVKDTADE IDLEPVAHSA LRGLLDVQRT VRDAPTGLDV AKLARITVIG E ADEARAAI RAWIAQAVTW HDPTMLGVAL AAPDLESGDW SWLKWLPHVD VPNEADGVGP ARYLTTSTAE LRERLAPALA DR PLFPAES GAALKHLLVV LDDPDADPDD IARKPGLTGV TVIHRTTELP NREQYPDPER PILRVADGRI ERWQVGGWQP CVD VADAMS AAEAAHIARR LSRWDSN |
-Macromolecule #4: ESX-3 secretion system protein EccE3
| Macromolecule | Name: ESX-3 secretion system protein EccE3 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria) Mycobacterium smegmatis (strain ATCC 700084 / mc(2)155) (bacteria)Strain: ATCC 700084 / mc(2)155 |
| Molecular weight | Theoretical: 30.850352 KDa |
| Recombinant expression | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Sequence | String: TARIALASLF VVAAVLAQPW QTTTQRWVLG VSIAAVIVLL AWWKGMFLTT RIGRALAMVR RNRAEDTVET DAHRATVVLR VDPAAPAQL PVVVGYLDRY GITCDKVRIT HRDAGGTRRS WISLTVDAVD NLAALQARSA RIPLQDTTEV VGRRLADHLR E QGWTVTVV ...String: TARIALASLF VVAAVLAQPW QTTTQRWVLG VSIAAVIVLL AWWKGMFLTT RIGRALAMVR RNRAEDTVET DAHRATVVLR VDPAAPAQL PVVVGYLDRY GITCDKVRIT HRDAGGTRRS WISLTVDAVD NLAALQARSA RIPLQDTTEV VGRRLADHLR E QGWTVTVV EGVDTPLPVS GKETWRGVAD DAGVVAAYRV KVDDRLDEVL AEIGHLPAEE TWTALEFTGS PAEPLLTVCA AV RTSDRPA AKAPLAGLTP ARGRHRPALA ALNPLSTERL DGTAVPLPA UniProtKB: ESX-3 secretion system protein EccE3 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL |
|---|---|
| Buffer | pH: 8 / Details: 30 mM Hepes pH 8.0, 150 mM NaCl |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK IV / Details: Blotting time: 3s Blotting force: -10. |
| Details | bound to Amphipol A8-35 |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number real images: 11903 / Average electron dose: 50.0 e/Å2 Details: Images acquired as 55 frames movies at a calibrated magnification of 1.0635 Angs/px |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.6 µm / Nominal defocus min: 1.6 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Overall B value: 29.4 / Target criteria: Map correlation coefficient |
|---|---|
| Output model |  PDB-6sgz: |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)