+Search query
-Structure paper
| Title | Architectures of Lipid Transport Systems for the Bacterial Outer Membrane. |
|---|---|
| Journal, issue, pages | Cell, Vol. 169, Issue 2, Page 273-285.e17, Year 2017 |
| Publish date | Apr 6, 2017 |
 Authors Authors | Damian C Ekiert / Gira Bhabha / Georgia L Isom / Garrett Greenan / Sergey Ovchinnikov / Ian R Henderson / Jeffery S Cox / Ronald D Vale /   |
| PubMed Abstract | How phospholipids are trafficked between the bacterial inner and outer membranes through the hydrophilic space of the periplasm is not known. We report that members of the mammalian cell entry (MCE) ...How phospholipids are trafficked between the bacterial inner and outer membranes through the hydrophilic space of the periplasm is not known. We report that members of the mammalian cell entry (MCE) protein family form hexameric assemblies with a central channel capable of mediating lipid transport. The E. coli MCE protein, MlaD, forms a ring associated with an ABC transporter complex in the inner membrane. A soluble lipid-binding protein, MlaC, ferries lipids between MlaD and an outer membrane protein complex. In contrast, EM structures of two other E. coli MCE proteins show that YebT forms an elongated tube consisting of seven stacked MCE rings, and PqiB adopts a syringe-like architecture. Both YebT and PqiB create channels of sufficient length to span the periplasmic space. This work reveals diverse architectures of highly conserved protein-based channels implicated in the transport of lipids between the membranes of bacteria and some eukaryotic organelles. |
 External links External links |  Cell / Cell /  PubMed:28388411 / PubMed:28388411 /  PubMed Central PubMed Central |
| Methods | EM (single particle) / X-ray diffraction |
| Resolution | 1.501 - 25.0 Å |
| Structure data | EMDB-8608, PDB-5uvn:  EMDB-8610:  EMDB-8611: 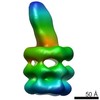 EMDB-8612:  PDB-5uw2:  PDB-5uw8:  PDB-5uwa:  PDB-5uwb: |
| Chemicals |  ChemComp-ZN:  ChemComp-HOH:  ChemComp-8ND:  ChemComp-PEF: |
| Source |
|
 Keywords Keywords | TRANSPORT PROTEIN / MCE protein / bacterial lipid transport / phospholipid-binding protein / Lipid-binding / periplasmic / MlaC / transport |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN Papers
About EMN Papers







