-検索条件
-検索結果
検索 (著者・登録者: white & ki)の結果208件中、1から50件目までを表示しています

EMDB-44635: 
Inactive mu opioid receptor bound to Nb6, naloxone and NAM

PDB-9bjk: 
Inactive mu opioid receptor bound to Nb6, naloxone and NAM
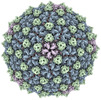
EMDB-40271: 
Mycobacterium phage Adjutor

PDB-8saj: 
Mycobacterium phage Adjutor

EMDB-42681: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the upper strand

EMDB-42682: 
The structure of the native cardiac thin filament troponin core in Ca2+-free tilted state from the upper strand

EMDB-42683: 
The structure of the native cardiac thin filament troponin core in Ca2+-free rotated state from the upper strand

EMDB-42800: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the lower strand

EMDB-42833: 
The structure of the native cardiac thin filament troponin core in Ca2+-free rotated state from the lower strand

EMDB-42835: 
The structure of the native cardiac thin filament troponin core in Ca2+-free tilted state from the lower strand

EMDB-42846: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state from the upper strand

EMDB-42847: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound partially activated state from the upper strand

EMDB-42849: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state 1 from the lower strand

EMDB-42856: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state 2 from the lower strand

EMDB-42858: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound partially activated state from the lower strand

EMDB-42874: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the upper strand activated by the C1-domain of cardiac myosin binding protein C

PDB-8uww: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the upper strand

PDB-8uwx: 
The structure of the native cardiac thin filament troponin core in Ca2+-free tilted state from the upper strand

PDB-8uwy: 
The structure of the native cardiac thin filament troponin core in Ca2+-free rotated state from the upper strand

PDB-8uyd: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the lower strand

PDB-8uz5: 
The structure of the native cardiac thin filament troponin core in Ca2+-free rotated state from the lower strand

PDB-8uz6: 
The structure of the native cardiac thin filament troponin core in Ca2+-free tilted state from the lower strand

PDB-8uzx: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state from the upper strand

PDB-8uzy: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound partially activated state from the upper strand

PDB-8v01: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state 1 from the lower strand

PDB-8v0i: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound fully activated state 2 from the lower strand

PDB-8v0k: 
The structure of the native cardiac thin filament troponin core in Ca2+-bound partially activated state from the lower strand

PDB-8v0y: 
The structure of the native cardiac thin filament troponin core in Ca2+-free state from the upper strand activated by the C1-domain of cardiac myosin binding protein C
EMDB-28198: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with LLNL-199

EMDB-28199: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with 2130-1-0114-112

PDB-8ekd: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with 2130-1-0114-112

EMDB-43013: 
Myxococcus xanthus HEnc-K417N(A) protein shell with icosahedral T=1 symmetry

EMDB-43016: 
Myxococcus xanthus HEnc-K417N(A) protein shell with tetrahedral symmetry (12 pentamers, 4 hexamers)

EMDB-43037: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D3 symmetry (12 pentamers, 3 hexamers)

EMDB-43038: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D6 symmetry (12 pentamers, 8 hexamers)

EMDB-43039: 
Myxococcus xanthus HEnc-K417N(A) protein shell with C2 symmetry (12 pentamers, 9 hexamers)

EMDB-43040: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D3 symmetry (12 pentamers, 11 hexamers)

EMDB-43041: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D2 symmetry (12 pentamers, 12 hexamers)

EMDB-43042: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D2 symmetry (12 pentamers, 14 hexamers)

EMDB-43043: 
Myxococcus xanthus HEnc-K417N(A) protein shell with D5 symmetry (12 pentamers, 15 hexamers)

EMDB-43113: 
Myxococcus xanthus EncA WT protein shell with icosahedral symmetry T=3

EMDB-43036: 
Myxococcus xanthus HEnc-K417N(A) protein shell with icosahedral T=3 symmetry

EMDB-40711: 
CryoEM structure of Western equine encephalitis virus VLP in complex with the chimeric Du-D1-Mo-D2 MXRA8 receptor

PDB-8sqn: 
CryoEM structure of Western equine encephalitis virus VLP in complex with the chimeric Du-D1-Mo-D2 MXRA8 receptor

EMDB-27272: 
CryoEM structure of Western equine encephalitis virus VLP
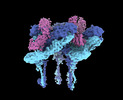
EMDB-27271: 
CryoEM structure of Western equine encephalitis virus VLP in complex with the avian MXRA8 receptor

EMDB-28761: 
Mycobacterium phage Che8 mutant capsid (gene 110 deletion)

EMDB-28644: 
CryoEM structure of Western equine encephalitis virus VLP in complex with the avian MXRA8 receptor

EMDB-29530: 
SARS-CoV-2 XBB.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-29531: 
SARS-CoV-2 BQ.1.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
