+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mycobacterium phage Che8 mutant capsid (gene 110 deletion) | |||||||||
 Map data Map data | Sharpened map of Che_mutant_map.mrc. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HK97-fold / T=9 / tailed bacteriophage / VIRUS | |||||||||
| Biological species |  Mycobacterium phage Che8 (virus) Mycobacterium phage Che8 (virus) | |||||||||
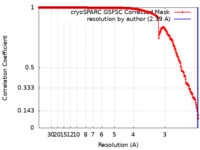
| Method | single particle reconstruction / cryo EM / Resolution: 2.39 Å | |||||||||
 Authors Authors | Podgorski JM / White SJ | |||||||||
| Funding support | 1 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Diverse O-linked Glycosylation of phage structural proteins is mediated through phage-encoded glycosyltransferases. Authors: Pope WH / Hatfull GF | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28761.map.gz emd_28761.map.gz | 1.5 GB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28761-v30.xml emd-28761-v30.xml emd-28761.xml emd-28761.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_28761_fsc.xml emd_28761_fsc.xml | 26.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_28761.png emd_28761.png | 316.3 KB | ||
| Masks |  emd_28761_msk_1.map emd_28761_msk_1.map | 1.9 GB |  Mask map Mask map | |
| Filedesc metadata |  emd-28761.cif.gz emd-28761.cif.gz | 4.2 KB | ||
| Others |  emd_28761_additional_1.map.gz emd_28761_additional_1.map.gz emd_28761_half_map_1.map.gz emd_28761_half_map_1.map.gz emd_28761_half_map_2.map.gz emd_28761_half_map_2.map.gz | 1.8 GB 1.8 GB 1.8 GB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28761 http://ftp.pdbj.org/pub/emdb/structures/EMD-28761 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28761 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28761 | HTTPS FTP |
-Related structure data
| Related structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28761.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28761.map.gz / Format: CCP4 / Size: 1.9 GB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened map of Che_mutant_map.mrc. | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.182 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_28761_msk_1.map emd_28761_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
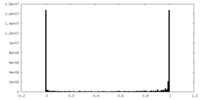
| Density Histograms |
-Additional map: Ewald sphere corrected map.
| File | emd_28761_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Ewald sphere corrected map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map of ewald sphere corrected Che mutant map.mrc.
| File | emd_28761_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of ewald sphere corrected Che_mutant_map.mrc. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map of ewald sphere corrected Che mutant map.mrc.
| File | emd_28761_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map of ewald sphere corrected Che_mutant_map.mrc. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Mycobacterium phage Che8
| Entire | Name:  Mycobacterium phage Che8 (virus) Mycobacterium phage Che8 (virus) |
|---|---|
| Components |
|
-Supramolecule #1: Mycobacterium phage Che8
| Supramolecule | Name: Mycobacterium phage Che8 / type: virus / ID: 1 / Parent: 0 / Details: MUTANT. Gene 110 deleted. / NCBI-ID: 2907829 / Sci species name: Mycobacterium phage Che8 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Mycolicibacterium smegmatis MC2 155 (bacteria) Mycolicibacterium smegmatis MC2 155 (bacteria) |
| Virus shell | Shell ID: 1 / Diameter: 730.0 Å / T number (triangulation number): 9 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 283 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 8118 / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)