-Search query
-Search result
Showing all 38 items for (author: weidle & c)
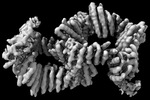
EMDB-42542: 
CryoEM Structure of Allosterically Switchable De Novo Protein sr322, In Closed State without Effector Peptide, off Target Multimeric State

PDB-8utm: 
CryoEM Structure of Allosterically Switchable De Novo Protein sr322, In Closed State without Effector Peptide, off Target Multimeric State

EMDB-42442: 
CryoEM Structure of Allosterically Switchable De Novo Protein sr322, In Closed State without Effector Peptide

PDB-8up1: 
CryoEM Structure of Allosterically Switchable De Novo Protein sr322, In Closed State without Effector Peptide
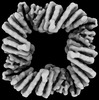
EMDB-42491: 
CryoEM Structure of Allosterically Switchable De Novo Protein sr312, in Open State with Effector Peptide

PDB-8ure: 
CryoEM Structure of Allosterically Switchable De Novo Protein sr312, in Open State with Effector Peptide

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
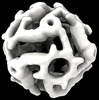
EMDB-42031: 
Computational Designed Nanocage O43_129_+8

EMDB-43318: 
Twistless helix 12 repeat ring design R12B

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage

EMDB-42906: 
Computational Designed Nanocage O43_129

EMDB-42944: 
Computational Designed Nanocage O43_129_+4

PDB-8gel: 
Cryo-EM structure of synthetic tetrameric building block sC4

PDB-8tl7: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage

PDB-8v2d: 
Computational Designed Nanocage O43_129

PDB-8v3b: 
Computational Designed Nanocage O43_129_+4

EMDB-27031: 
Accurate computational design of genetically encoded 3D protein crystals

EMDB-40926: 
CryoEM Structure of Computationally Designed Nanocage O32-ZL4

PDB-8cwy: 
Accurate computational design of genetically encoded 3D protein crystals

PDB-8szz: 
CryoEM Structure of Computationally Designed Nanocage O32-ZL4

EMDB-29915: 
CryoEM map of a de novo designed octahedral nanocage with programmable volume; design cage_O4_34

EMDB-40070: 
Cryo-EM map of synthetic cage_O3_10 reconstructed without symmetry (C1)

EMDB-40071: 
Cryo-EM map of synthetic cage_O3_10 reconstructed with O symmetry

EMDB-40073: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1), with 1 monomer missing (class 3.0)

EMDB-40074: 
Cryo-EM map of synthetic cage_T3_5 reconstructed with T symmetry

EMDB-40075: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1)

EMDB-40076: 
Cryo-EM map of synthetic cage_T3_5+2 reconstructed without symmetry (C1)

EMDB-40072: 
Cryo-EM map of synthetic cage_T3_5 reconstructed without symmetry (C1), with 1 trimer missing (class 3.1)

EMDB-9294: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound, sharpened map

EMDB-9295: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound, unsharpened map

EMDB-9303: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 2 Fabs bound, sharpened map

EMDB-9304: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 2 Fabs bound, unsharpened map

PDB-6myy: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound, sharpened map

PDB-6mzj: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 2 Fabs bound, sharpened map

EMDB-7344: 
An anti-gH/gL antibody that neutralizes dual-tropic infection defines a site of vulnerability on Epstein-Barr virus

EMDB-7345: 
An anti-gH/gL antibody that neutralizes dual-tropic infection defines a site of vulnerability on Epstein-Barr virus

PDB-6c5v: 
An anti-gH/gL antibody that neutralizes dual-tropic infection defines a site of vulnerability on Epstein-Barr virus
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
