-検索条件
-検索結果
検索 (著者・登録者: wang & h)の結果10,205件中、1から50件目までを表示しています

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
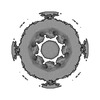
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
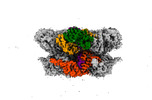
EMDB-42603: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/hexamer)
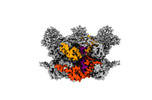
EMDB-42625: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC804515)
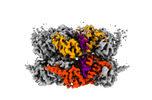
EMDB-42626: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/up)
EMDB-42627: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/down)
EMDB-44748: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/dodecamer)

PDB-8uv2: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/hexamer)

PDB-8uvo: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC804515)

PDB-8uvp: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/up)

PDB-8uvq: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/down)

PDB-9boq: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/dodecamer)

EMDB-60410: 
Cryo-EM structure of human ZnT1, in the absence of zinc, determined in outward-facing conformation

EMDB-60435: 
Cryo-EM structure of human ZnT1, in the presence of zinc, determined in outward-facing conformation

EMDB-60099: 
SARS-CoV-2 spike trimer (6P) in complex with two R1-26 Fabs

EMDB-60100: 
SARS-CoV-2 spike trimer (6P) in complex with three R1-26 Fabs

EMDB-60101: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, head-to-head aggregate

EMDB-60102: 
SARS-CoV-2 spike trimer (6P) in complex with R1-26 Fab, focused refinement of RBD-Fab region

EMDB-60103: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 Fabs

EMDB-60104: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs

EMDB-60105: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C1 symmetry)

EMDB-60106: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 Fabs, head-to-head aggregate (C3 symmetry)

EMDB-60107: 
SARS-CoV-2 spike trimer (6P) in complex with two H18 and two R1-32 Fabs

EMDB-60108: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs

EMDB-60109: 
SARS-CoV-2 spike trimer (6P) in complex with three H18 and three R1-32 Fabs (one RBD rotated)

EMDB-60110: 
SARS-CoV-2 S1 in complex with H18 and R1-32 Fab

EMDB-60111: 
Dimer of SARS-CoV-2 S1 in complex with H18 and R1-32 Fabs
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
