-Search query
-Search result
Showing 1 - 50 of 2,704 items for (author: thomas & g)

EMDB-45001: 
Structure of Mnx H340A complex from Bacillus sp. PL-12

PDB-9bxa: 
Structure of Mnx H340A complex from Bacillus sp. PL-12

EMDB-18779: 
Structure of the non-mitochondrial citrate synthase from Ananas comosus

PDB-8qzp: 
Structure of the non-mitochondrial citrate synthase from Ananas comosus

EMDB-18990: 
CryoEM map of tau PHF sarkosyl-extracted from a human AD patient (associated with in situ tomography)

EMDB-50148: 
Tau PHF subtomogram average relating to CS1 extended data Figure 9A

EMDB-50152: 
Tau PHF subtomogram average relating to CS2 Figure 3i-j.

EMDB-50153: 
Tau PHF subtomogram average relating to CS3 extended data Figure 9c

EMDB-50155: 
Tau PHF subtomogram average relating to CS4 extended data Figure 9d

EMDB-50156: 
Tau PHF subtomogram average relating to CS5 extended data Figure 9b

EMDB-50157: 
Tau PHF subtomogram average relating to CS6 extended data Figure 9e

EMDB-50159: 
Tau PHF subtomogram average relating to CS7 extended data Figure 9f

EMDB-50160: 
Tau PHF subtomogram average relating to LOL1_PHF Figure 4g-h

EMDB-50161: 
Tau SF subtomogram average relating to LOL1_SF Figure 4g-h

EMDB-50162: 
Tau SF subtomogram average relating to LOL2_SF Figure 4i-j

EMDB-19846: 
PHF type tau filament from V337M mutant

EMDB-19849: 
PHF type tau filament from V337M mutant

EMDB-19852: 
PHF type tau filament from V337M mutant
PDB-9eo7: 
PHF type tau filament from V337M mutant

PDB-9eo9: 
SF type tau filament from V337M mutant
PDB-9eoe: 
TF type tau filament from V337M mutant
EMDB-50358: 
In vitro-induced genome-releasing intermediate of Rhodobacter microvirus Ebor computed with C5 symmetry

EMDB-19929: 
Structural basis of D9-THC analog activity at the Cannabinoid 1 receptor

PDB-9erx: 
Structural basis of D9-THC analog activity at the Cannabinoid 1 receptor

EMDB-40046: 
CryoEM structure of Influenza A virus A/Melbourner/1/1946 (H1N1) hemagglutinin bound to GS10-X6-BE4 Fab

PDB-8ghk: 
CryoEM structure of Influenza A virus A/Melbourner/1/1946 (H1N1) hemagglutinin bound to GS10-X6-BE4 Fab

EMDB-19163: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.

EMDB-19164: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.

EMDB-19165: 
Trimeric HSV-2F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.

EMDB-19166: 
Trimeric HSV-2G gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.

PDB-8rgz: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.

PDB-8rh0: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.

PDB-8rh1: 
Trimeric HSV-2F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.

PDB-8rh2: 
Trimeric HSV-2G gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.

EMDB-16375: 
SARS-CoV2 Omicron BA.1 RBD in complex with CAB-A17 antibody

PDB-8c0y: 
SARS-CoV2 Omicron BA.1 RBD in complex with CAB-A17 antibody

EMDB-50356: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry

EMDB-50357: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
EMDB-50359: 
Rhodobacter microvirus Ebor attached to B10 host cell reconstructed by single particle analysis with applied C5 symmetry
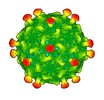
EMDB-50360: 
Rhodobacter microvirus Ebor attached to the outer membrane vesicle

EMDB-50361: 
Rhodobacter microvirus Ebor attached to the host cell reconstructed by subtomogram averaging

PDB-9ffg: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry

PDB-9ffh: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry

EMDB-28966: 
CryoEM map of de novo designed oligomeric protein C4-71_6x

EMDB-28967: 
CryoEM map of de novo designed oligomeric protein C4-71_8x
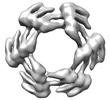
EMDB-28968: 
CryoEM map of de novo designed oligomeric protein C6-71

EMDB-28969: 
CryoEM map of de novo designed oligomeric protein C6-71_6x

EMDB-28970: 
CryoEM map of de novo designed oligomeric protein C6-71_8x

EMDB-28971: 
CryoEM map of de novo designed oligomeric protein C8-71_6x

EMDB-28972: 
CryoEM map of de novo designed oligomeric protein C8-71_8x
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
