-Search query
-Search result
Showing 1 - 50 of 646 items for (author: smith & am)

EMDB-17295: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

PDB-8oyt: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

EMDB-40825: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

PDB-8sx3: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

EMDB-17296: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '2 up 1 down' RBD conformation

PDB-8oyu: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '2 up 1 down' RBD conformation

EMDB-41024: 
MD65 N332-GT5 SOSIP in complex with RM_N332_03 Fab and RM20A3 Fab

EMDB-41025: 
MD65 N332-GT5 SOSIP in complex with RM_N332_36 Fab and RM20A3 Fab

EMDB-41026: 
MD65 N332-GT5 SOSIP in complex with RM_N332_32 Fab and RM20A3

EMDB-41027: 
MD65 N332-GT5 SOSIP in complex with RM_N332_08 Fab and RM20A3 Fab

EMDB-41034: 
MD64 N332-GT5 SOSIP

EMDB-41035: 
MD65 N332-GT5 SOSIP in complex with RM_N332_07 Fab and RM20A3 Fab

PDB-8t49: 
MD65 N332-GT5 SOSIP in complex with RM_N332_03 Fab and RM20A3 Fab

PDB-8t4a: 
MD65 N332-GT5 SOSIP in complex with RM_N332_36 Fab and RM20A3 Fab

PDB-8t4b: 
MD65 N332-GT5 SOSIP in complex with RM_N332_32 Fab and RM20A3

PDB-8t4d: 
MD65 N332-GT5 SOSIP in complex with RM_N332_08 Fab and RM20A3 Fab

PDB-8t4k: 
MD64 N332-GT5 SOSIP

PDB-8t4l: 
MD65 N332-GT5 SOSIP in complex with RM_N332_07 Fab and RM20A3 Fab

EMDB-41933: 
CryoEM map of horse spleen apoferritin determined as a reference for benchmarking square and rectangular apertures for cryo-EM

EMDB-41936: 
CryoEM map of horse spleen apoferritin determined as a reference for benchmarking square and rectangular apertures for cryo-EM (Falcon IV round beam)

EMDB-41937: 
CryoEM map of horse spleen apoferritin determined for benchmarking square and rectangular apertures for cryo-EM (Falcon IV square beam)

EMDB-41938: 
CryoEM map of horse spleen apoferritin determined for benchmarking square and rectangular apertures for cryo-EM (Gatan K3 rectangular beam)

EMDB-42882: 
Amylin 1 receptor bound to salmon calcitonin (Amy1R:sCT) reconstructed from cryo-EM datasets recorded using a rectangular aperture
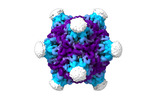
EMDB-19024: 
Structure of the PNMA2 capsid
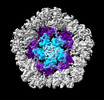
EMDB-19025: 
Structure of the five-fold capsomer of the PNMA2 capsid
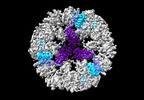
EMDB-19026: 
Structure of the three-fold capsomer of the PNMA2 capsid
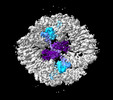
EMDB-19027: 
Structure of the two-fold capsomer of the PNMA2 capsid

PDB-8rb3: 
Structure of the PNMA2 capsid

PDB-8rb4: 
Structure of the five-fold capsomer of the PNMA2 capsid

PDB-8rb5: 
Structure of the three-fold capsomer of the PNMA2 capsid

PDB-8rb7: 
Structure of the two-fold capsomer of the PNMA2 capsid

EMDB-42600: 
Murine norovirus in the presence of 1mM calcium

EMDB-42604: 
Murine norovirus + 1 mM MgCl2

EMDB-42623: 
Murine norovirus dialyzed against EDTA

PDB-8uux: 
Murine norovirus capsid protein in the presence of 1mM calcium

PDB-8uv3: 
Murine norovirus capsid protein + 1 mM MgCl2

EMDB-29036: 
Amyloid-beta (1-40) fibrils derived from a CAA patient

EMDB-29037: 
Amyloid-beta (1-40) fibrils derived from familial Dutch-type CAA patient (population B)

EMDB-29038: 
Amyloid-beta (1-40) fibrils derived from familial Dutch-type CAA patient (population A)

PDB-8ff2: 
Amyloid-beta (1-40) fibrils derived from a CAA patient

PDB-8ff3: 
Amyloid-beta (1-40) fibrils derived from familial Dutch-type CAA patient (population B)

EMDB-16328: 
Outer membrane attachment porin OmpM1 from Veillonella parvula

EMDB-16332: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, native

EMDB-16333: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, C3 symmetry

PDB-8bym: 
Outer membrane attachment porin OmpM1 from Veillonella parvula

PDB-8bys: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, native

PDB-8byt: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, C3 symmetry

EMDB-40464: 
CCT G beta 5 complex intermediate state

EMDB-40486: 
CCT-G beta 5 complex closed state 7

PDB-8sgq: 
CCT G beta 5 complex intermediate state
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
