-Search query
-Search result
Showing 1 - 50 of 539 items for (author: singh & j)
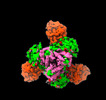
EMDB-36815: 
Recognition determinants of broad and potent HIV-1 neutralization by an affinity matured antibody from a pediatric elite-neutralizer
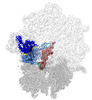
EMDB-40929: 
E. coli initiation complex with EQ2-EttA in Hydrolytic 2 conformation
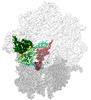
EMDB-40928: 
E. coli initiation complex with EQ2-YbiT in Non-hydrolytic 2/PtIM(b) conformation

EMDB-40921: 
E. coli pre-elongation complex without an A-site tRNA with EQ2-EttA in Hydrolytic 1 conformation
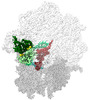
EMDB-40924: 
E. coli initiation complex with EQ2-YbiT in Hydrolytic 2/PtIM(a) conformation
EMDB-40925: 
E. coli initiation complex with EQ2-YbiT in Intermediate/PtIM(b) conformation
EMDB-40927: 
E. coli initiation complex with EQ2-YbiT in Non-hydrolytic 1/PtIM(a) conformation

EMDB-40932: 
E. coli 70S initiation complex

EMDB-40923: 
E. coli pre-elongation complex without an A-site tRNA with EQ2-YbiT in Non-hydrolytic 1/PtIM(a) conformation

EMDB-40939: 
E. coli 70S initiation complex (bL33 absent)
EMDB-34548: 
Spike 3-up RBD with THSC20.HVTR04 (Fab4): State - III
EMDB-34563: 
Spike 2-up RBD with THSC20.HVTR26 (Fab26): State - I

EMDB-34602: 
State - I: Spike 2-up RBD with THSC20.HVTR26 (Fab26)
EMDB-34603: 
State - II: Spike 3-up RBD with THSC20.HVTR26 (Fab26)
EMDB-34546: 
State - I: Spike 2-up RBD with THSC20.HVTR04 (Fab4)
EMDB-34547: 
Spike 3-up RBD with THSC20.HVTR04 (Fab4): State - II

EMDB-17835: 
Consensus cryo-EM structure of Dynein-Dynactin-JIP3(1-185)-LIS1

EMDB-17836: 
Consensus cryo-EM structure of Dynein-dynactin-JIP3(1-560)-LIS1

EMDB-17825: 
Cytoplasmic dynein-1 motor domain in post-powerstroke state

EMDB-17826: 
Cytoplasmic dynein-1 motor domain bound to dynactin-p150glued and LIS1

EMDB-17828: 
Cytoplasmic dynein-1 motor domain bound to LIS1

EMDB-17829: 
Cytoplasmic dynein-1 A1/A2 motor domains bound to LIS1

EMDB-17830: 
Cytoplasmic dynein-A heavy chain bound to dynactin-p150glued and IC-LC tower

EMDB-17831: 
Cytoplasmic dynein-B heavy chain bound to IC-LC tower

EMDB-17832: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-LZI

EMDB-17833: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-RH1

EMDB-17834: 
Dynactin pointed end bound to JIP3

EMDB-17873: 
Composite structure of Dynein-Dynactin-JIP3-LIS1

PDB-8pqv: 
Cytoplasmic dynein-1 motor domain in post-powerstroke state

PDB-8pqw: 
Cytoplasmic dynein-1 motor domain bound to dynactin-p150glued and LIS1

PDB-8pqy: 
Cytoplasmic dynein-1 motor domain bound to LIS1

PDB-8pqz: 
Cytoplasmic dynein-1 A1/A2 motor domains bound to LIS1

PDB-8pr0: 
Cytoplasmic dynein-A heavy chain bound to dynactin-p150glued and IC-LC tower

PDB-8pr1: 
Cytoplasmic dynein-B heavy chain bound to IC-LC tower

PDB-8pr2: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-LZI

PDB-8pr3: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-RH1

PDB-8pr4: 
Dynactin pointed end bound to JIP3

PDB-8pr5: 
Structure of the autoinhibited dynactin p150glued projection

PDB-8ptk: 
Composite structure of Dynein-Dynactin-JIP3-LIS1

EMDB-42628: 
Structure of the insect gustatory receptor Gr9 from Bombyx mori

EMDB-42629: 
Structure of the insect gustatory receptor Gr9 from Bombyx mori in complex with D-fructose

EMDB-43548: 
Structure of the insect gustatory receptor Gr9 from Bombyx mori in complex with L-sorbose

PDB-8uvt: 
Structure of the insect gustatory receptor Gr9 from Bombyx mori

PDB-8uvu: 
Structure of the insect gustatory receptor Gr9 from Bombyx mori in complex with D-fructose

PDB-8vv3: 
Structure of the insect gustatory receptor Gr9 from Bombyx mori in complex with L-sorbose

EMDB-35817: 
Structure of Niacin-GPR109A-G protein complex

EMDB-35822: 
Structure of MK6892-GPR109A-G-protein complex

EMDB-35831: 
Structure of GSK256073-GPR109A-G-protein complex

EMDB-36193: 
Structure of Acipimox-GPR109A-G protein complex

EMDB-36280: 
Structure of MMF-GPR109A-G protein complex
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
