-Search query
-Search result
Showing 1 - 50 of 92 items for (author: robb & c)

EMDB-43753: 
Yeast U1 snRNP with humanized U1C Zinc-Finger domain

PDB-8w2o: 
Yeast U1 snRNP with humanized U1C Zinc-Finger domain
EMDB-42464: 
chEnv TTT protein in complex with 43A2 Fab

EMDB-42468: 
chEnv TTT protein in complex with CM01A Fab
EMDB-43664: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines
EMDB-43665: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines (cH125 TTT)

EMDB-43666: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines (H2/1 GCN4)
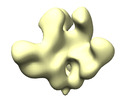

EMDB-43668: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines (H5/1 GCN4)
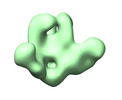

EMDB-43669: 
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines. H5 GCN4

EMDB-26560: 
Cryo-EM structure of SARS-CoV-2 Alpha (B.1.1.7) spike protein in complex with bebtelovimab

EMDB-28233: 
Cryo-EM structure of LRP2 at pH 7.5

EMDB-28241: 
Cryo-EM structure of LRP2 at pH 5.2

EMDB-28242: 
Local refinement map of LRP2 P1-P2 domains at pH 7.5

EMDB-28243: 
Local refinement of P3-P6 domains of LRP2 at pH 7.5

EMDB-28250: 
Local refinement of the P7 domain of LRP2 at pH 7.5

EMDB-28251: 
Local refinement of the R4 domain of LRP2 at pH 7.5

EMDB-28252: 
Local refinement of P8 domain of LRP2 at pH 7.5

EMDB-28253: 
Local refinement of P1-P2 domains of LRP2 at pH 5.2

EMDB-28258: 
Local refinement of P3-P6 domains of LRP2 at pH 5.2

EMDB-28260: 
Local refinement of P7 domain of LRP2 at pH 5.2

EMDB-28261: 
Local refinement of R3 domain of LRP2 at pH 5.2

EMDB-28265: 
Local refinement of P8 domain of LRP2 at pH 5.2

PDB-8em4: 
Cryo-EM structure of LRP2 at pH 7.5

PDB-8em7: 
Cryo-EM structure of LRP2 at pH 5.2

EMDB-27177: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein

PDB-8d48: 
sd1.040 Fab in complex with SARS-CoV-2 Spike 2P glycoprotein

EMDB-13594: 
Cryo-EM structure of Saccharomyces cerevisiae TOROID (TORC1 Organized in Inhibited Domains).

EMDB-13595: 
Helical reconstruction of TOROID (TORC1 Organized in Inhibited Domains) filaments.

PDB-7pqh: 
Cryo-EM structure of Saccharomyces cerevisiae TOROID (TORC1 Organized in Inhibited Domains).

EMDB-15364: 
Cryo-EM structure of the SEA complex (consensus map)

EMDB-15373: 
Cryo-EM structure of the SEA complex (protomer focused map)

EMDB-15374: 
Cryo-EM structure of the SEA complex (Sea2-Sea3 focused map)

EMDB-15381: 
Cryo-EM structure of the SEA complex (wing focused map)

PDB-8adl: 
Cryo-EM structure of the SEA complex

PDB-8ae6: 
Cryo-EM structure of the SEA complex wing (SEACIT)

EMDB-25107: 
Lassa virus glycoprotein construct(Josiah GPCysR4) recovered from GPC-I53-50 nanoparticle by localized reconstruction

EMDB-25108: 
I53-50 nanoparticle core reconstructed from GPC-I53-50NP by focused refinement

EMDB-25109: 
Lassa virus glycoprotein construct (Josiah GPC-I53-50A) in complex with LAVA01 antibody

PDB-7sgd: 
Lassa virus glycoprotein construct(Josiah GPCysR4) recovered from GPC-I53-50 nanoparticle by localized reconstruction

PDB-7sge: 
I53-50 nanoparticle core reconstructed from GPC-I53-50NP by focused refinement

PDB-7sgf: 
Lassa virus glycoprotein construct (Josiah GPC-I53-50A) in complex with LAVA01 antibody
EMDB-26217: 
Negative stain EM map of COVA1-07 mAb bound to the S2 domain of SARS-CoV-2 S
EMDB-26218: 
Negative stain EM map of COVA2-14 mAb bound to the S2 domain of SARS-CoV-2 S

EMDB-26219: 
Negative stain EM map of COVA2-18 mAb bound to the S2 domain of SARS-CoV-2 S
EMDB-26220: 
Negative stain EM map of the S2 domain of SARS-CoV-2 S

EMDB-13776: 
Structure of formaldehyde cross-linked SARS-CoV-2 S glycoprotein

PDB-7q1z: 
Structure of formaldehyde cross-linked SARS-CoV-2 S glycoprotein

EMDB-11818: 
Cryo-EM structure of the divergent actomyosin complex from Plasmodium falciparum Myosin A in the Rigor state

PDB-7aln: 
Cryo-EM structure of the divergent actomyosin complex from Plasmodium falciparum Myosin A in the Rigor state

EMDB-23265: 
Computationally designed icosahedral antibody nanocage with Fc i52.3+Fc
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
