-Search query
-Search result
Showing 1 - 50 of 4,532 items for (author: michael & b)

EMDB-41903: 
Cryo-EM structure of PsBphP in Pr state

EMDB-41941: 
Cryo-EM structure of PsBphP in Pfr state, Dimer of Dimers FL

EMDB-41942: 
Cryo-EM structure of PsBphP in Pfr state, Dimer of Dimers PSM only

EMDB-41943: 
Cryo-EM structure of PsBphP in Pfr state, medial PSM only

EMDB-41944: 
Cryo-EM structure of PsBphP in Pfr state, splayed PSM only

EMDB-42030: 
Cryo-EM structure of PsBphP in Pr state, extended DHp

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
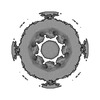
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map

EMDB-19209: 
TadA/CpaF with ADP

EMDB-19275: 
TadA/CpaF with AMPPNP

EMDB-19279: 
TadA/CpaF nucleotide free

PDB-8rjf: 
TadA/CpaF with ADP

PDB-8rkd: 
TadA/CpaF with AMPPNP

PDB-8rkl: 
TadA/CpaF nucleotide free

EMDB-43814: 
Cryo-EM structure of the active Lactococcus lactis Csm bound to target in post-cleavage stage

EMDB-43815: 
Cryo-EM structure of the active Lactococcus lactis Csm bound to target in pre-cleavage stage

PDB-9ash: 
Cryo-EM structure of the active Lactococcus lactis Csm bound to target in post-cleavage stage

PDB-9asi: 
Cryo-EM structure of the active Lactococcus lactis Csm bound to target in pre-cleavage stage
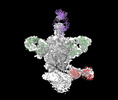
EMDB-44246: 
Cryo-EM structure of HIV-1 JRFL v6 Env in complex with vaccine-elicited, Membrane Proximal External Region (MPER) directed antibody DH1317.4.

EMDB-40218: 
CryoEM structure of TnsC(1-503) bound to TnsD(1-318) from E.coli Tn7

EMDB-40221: 
CryoEM structure of the TnsC(1-503)-TnsD(1-318)-DNA complex in a 7:2:1 stoichiometry from E. coli Tn7

EMDB-40222: 
CryoEM structure of the TnsC(1-503)-TnsD(1-318)-DNA complex in a 6:2:1 stoichiometry from E. coli Tn7

EMDB-43138: 
CryoEM structure of the TnsC(1-503)-TnsD(1-318)-DNA complex in a 7:2:1 stoichiometry from E. coli Tn7 bound to ATPgS and ADP

EMDB-43140: 
CyoEM structure of the TnsC(1-503)-TnsD(1-318)-DNA complex in a 6:2:1 stoichiometry from E. coli Tn7 bound to ATPgS and ADP

PDB-8glu: 
CryoEM structure of TnsC(1-503) bound to TnsD(1-318) from E.coli Tn7

PDB-8glw: 
CryoEM structure of the TnsC(1-503)-TnsD(1-318)-DNA complex in a 7:2:1 stoichiometry from E. coli Tn7

PDB-8glx: 
CryoEM structure of the TnsC(1-503)-TnsD(1-318)-DNA complex in a 6:2:1 stoichiometry from E. coli Tn7
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
