-Search query
-Search result
Showing 1 - 50 of 1,150 items for (author: mani & n)
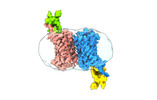
EMDB-36488: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Composite map)

EMDB-37212: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Receptor original map)
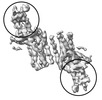
EMDB-37214: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Ligand/CCL7 focused map)

PDB-8jps: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Composite map)

EMDB-43762: 
Aca2 from Pectobacterium phage ZF40 bound to RNA

PDB-8w35: 
Aca2 from Pectobacterium phage ZF40 bound to RNA

EMDB-50672: 
A 3.3A sub-tomogram average of HIV-1 CA-SP1 from 5 tomograms in EMPIAR-10164 obtained using RELION 5

EMDB-42676: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

EMDB-42999: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

PDB-8uwl: 
5-HT2AR bound to Lisuride in complex with a mini-Gq protein and an active-state stabilizing single-chain variable fragment (scFv16) obtained by cryo-electron microscopy (cryoEM)

PDB-8v6u: 
5HT2AR-miniGq heterotrimer in complex with a novel agonist obtained from large scale docking

EMDB-35163: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 5.5

EMDB-35164: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in closed state
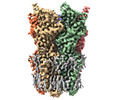
EMDB-36339: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 2.5
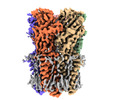
EMDB-37446: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in intermediate state
EMDB-37447: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in open state

PDB-8i47: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 5.5

PDB-8i48: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in closed state

PDB-8jj3: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 2.5

PDB-8wcq: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in intermediate state

PDB-8wcr: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 4 in open state

EMDB-35161: 
Cryo-EM structure of nanodisc (asolectin) reconstituted GLIC at pH 7.5

EMDB-35162: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 7.5

PDB-8i41: 
Cryo-EM structure of nanodisc (asolectin) reconstituted GLIC at pH 7.5

PDB-8i42: 
Cryo-EM structure of nanodisc (PE:PS:PC) reconstituted GLIC at pH 7.5

EMDB-17835: 
Consensus cryo-EM structure of Dynein-Dynactin-JIP3(1-185)-LIS1

EMDB-17836: 
Consensus cryo-EM structure of Dynein-dynactin-JIP3(1-560)-LIS1
EMDB-43647: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant
EMDB-43650: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant

EMDB-43656: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)

EMDB-43657: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant (Locally refined map)

PDB-8vy7: 
CryoEM structure of Gi-coupled TAS2R14 with cholesterol and an intracellular tastant

PDB-8vy9: 
CryoEM structure of Ggust-coupled TAS2R14 with cholesterol and an intracellular tastant

EMDB-17825: 
Cytoplasmic dynein-1 motor domain in post-powerstroke state

EMDB-17826: 
Cytoplasmic dynein-1 motor domain bound to dynactin-p150glued and LIS1

EMDB-17828: 
Cytoplasmic dynein-1 motor domain bound to LIS1

EMDB-17829: 
Cytoplasmic dynein-1 A1/A2 motor domains bound to LIS1

EMDB-17830: 
Cytoplasmic dynein-A heavy chain bound to dynactin-p150glued and IC-LC tower

EMDB-17831: 
Cytoplasmic dynein-B heavy chain bound to IC-LC tower

EMDB-17832: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-LZI

EMDB-17833: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-RH1

EMDB-17834: 
Dynactin pointed end bound to JIP3

EMDB-17873: 
Composite structure of Dynein-Dynactin-JIP3-LIS1

PDB-8pqv: 
Cytoplasmic dynein-1 motor domain in post-powerstroke state

PDB-8pqw: 
Cytoplasmic dynein-1 motor domain bound to dynactin-p150glued and LIS1

PDB-8pqy: 
Cytoplasmic dynein-1 motor domain bound to LIS1

PDB-8pqz: 
Cytoplasmic dynein-1 A1/A2 motor domains bound to LIS1

PDB-8pr0: 
Cytoplasmic dynein-A heavy chain bound to dynactin-p150glued and IC-LC tower

PDB-8pr1: 
Cytoplasmic dynein-B heavy chain bound to IC-LC tower

PDB-8pr2: 
Cytoplasmic dynein-1 heavy chain bound to JIP3-LZI
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
