-Search query
-Search result
Showing 1 - 50 of 763 items for (author: lo & hs)

EMDB-39212: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH8.0 (3.23A)

EMDB-39213: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH6.5 (2.82A)

EMDB-39214: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH5.0 (3.52A)

EMDB-39215: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH6.5 (3.12A)

EMDB-39217: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH5.0 (4.36A)

PDB-8yf6: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH8.0 (3.23A)

PDB-8yf7: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH6.5 (2.82A)

PDB-8yf8: 
Cryo-EM structure of Dragon Grouper nervous necrosis virus-like particle at pH5.0 (3.52A)

PDB-8yf9: 
Cryo-EM structure of Dragon Grouper nervous necrosis virion at pH6.5 (3.12A)
EMDB-42516: 
HIV-1 JR-FL NFL.664 soluble trimer in complex with polyclonal Fab from rabbit U5902

EMDB-42517: 
HIV-1 JR-FL NFL.664 soluble trimer in complex with polyclonal Fab from rabbit U5756

EMDB-42518: 
HIV-1 1086c NFL.664 soluble trimer in complex with polyclonal Fab from rabbit U5403

EMDB-42519: 
HIV-1 1086c NFL.664 soluble trimer in complex with polyclonal Fab from rabbit U5919

EMDB-19845: 
Outward-open structure of human dopamine transporter bound to cocaine

PDB-9eo4: 
Outward-open structure of human dopamine transporter bound to cocaine

EMDB-39126: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly

EMDB-39127: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly

PDB-8ybx: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly
EMDB-40248: 
CRISPR-Cas type III-D effector complex
EMDB-40250: 
CRISPR-Cas type III-D effector complex bound to a self-target RNA in the pre-cleavage state
EMDB-40251: 
CRISPR-Cas type III-D effector complex bound to self-target RNA in a post-cleavage state

EMDB-40276: 
CRISPR-Cas type III-D effector complex consensus map

EMDB-40296: 
CRISPR-Cas type III-D effector complex local refinement map

EMDB-40297: 
CRISPR-Cas type III-D effector complex bound to a target RNA local refinement map

EMDB-40298: 
CRISPR-Cas type III-D effector complex bound to a target RNA consensus map

PDB-8s9t: 
CRISPR-Cas type III-D effector complex

PDB-8s9v: 
CRISPR-Cas type III-D effector complex bound to a self-target RNA in the pre-cleavage state

PDB-8s9x: 
CRISPR-Cas type III-D effector complex bound to self-target RNA in a post-cleavage state

EMDB-40260: 
CryoEM map of a de novo designed T=4 icosahedral nanocage hierarchically built from pseudosymmetric trimers; design Ico(T=4)-4
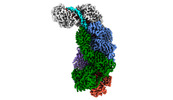
EMDB-40249: 
CRISPR-Cas type III-D effector complex bound to a target RNA

EMDB-18334: 
Cryo-EM structure of the inward-facing FLVCR1

EMDB-18335: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1

EMDB-18336: 
Cryo-EM structure of the inward-facing FLVCR2
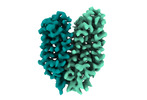
EMDB-18337: 
Cryo-EM structure of the outward-facing FLVCR2

EMDB-18339: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2

EMDB-19009: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1

PDB-8qcs: 
Cryo-EM structure of the inward-facing FLVCR1

PDB-8qct: 
Cryo-EM structure of the inward-facing choline-bound FLVCR1

PDB-8qcx: 
Cryo-EM structure of the inward-facing FLVCR2

PDB-8qcy: 
Cryo-EM structure of the outward-facing FLVCR2

PDB-8qd0: 
Cryo-EM structure of the inward-facing choline-bound FLVCR2

PDB-8r8t: 
Cryo-EM structure of the inward-facing ethanolamine-bound FLVCR1

EMDB-40267: 
CryoEM map of a T=1 off-target state of design Ico(T=4)-4

EMDB-40268: 
CryoEM map of a de novo designed T=4 octahedral nanocage hierarchically built from pseudosymmetric trimers; design Oct(T=4)-3

EMDB-40269: 
CryoEM map of a T=1 off-target state of design Oct(T=4)-3
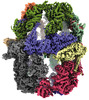
EMDB-19477: 
Saccharomyces cerevisiae FAS type I

EMDB-19489: 
Tobacco mosaic virus from scanning transmission electron microscopy at CSA=2.0 mrad
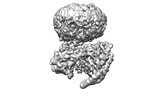
EMDB-42023: 
GPR3 Orphan G-coupled Protein Receptor in complex with Dominant Negative Gs.

PDB-8u8f: 
GPR3 Orphan G-coupled Protein Receptor in complex with Dominant Negative Gs.

EMDB-42480: 
Cryo-EM reconstruction of Staphylococcus aureus Oleate hydratase (OhyA) dimer with an ordered C-terminal membrane-association domain
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
