-Search query
-Search result
Showing all 42 items for (author: jones & ct)

PDB-7l6o: 
Cryo-EM structure of HIV-1 Env CH848.3.D0949.10.17chim.6R.DS.SOSIP.664
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23518: 
Cryo-EM map of DH851.3 bound to HIV-1 CH505 Env
Method: single particle / : Edwards RJ, Manne K, Acharya P

PDB-7lu9: 
Cryo-EM structure of DH851.3 bound to HIV-1 CH505 Env
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23519: 
Cryo-EM map of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Edwards RJ, Manne K, Acharya P

PDB-7lua: 
Cryo-EM structure of DH898.1 Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23152: 
Cryo-electron microscopy reconstruction of antibody DH898.1 Fab-dimer bound to glycans 332, 392, and 396 of HIV Env CH848 10.17 SOSIP trimer
Method: single particle / : Edwards RJ, Acharya P

EMDB-23153: 
Cryo-electron microscopy local refinement of antibody DH898.1 Fab-dimer bound to glycans 332, 392, and 396 of HIV Env CH848 10.17 SOSIP trimer
Method: single particle / : Edwards RJ, Acharya P

EMDB-23149: 
Cryo-electron microscopy reconstruction of antibody DH898.1 Fab-dimer bound near the CD4 binding site of HIV Env SOSIP trimer CH848 10.17
Method: single particle / : Edwards RJ, Acharya P

EMDB-23124: 
Cryo-electron microcospy reconstruction of CH848.3.D0949.10.17chim.6R.DS.SOSIP.664 HIV Env
Method: single particle / : Edwards RJ, Acharya P

EMDB-23145: 
Cryo-electron microscopy reconstruction of locally refined antibody DH898.1 Fab-dimer
Method: single particle / : Edwards RJ, Acharya P

PDB-7l6m: 
Cryo-EM structure of DH898.1 Fab-dimer from local refinement of the Fab-dimer bound near the CD4 binding site of HIV-1 Env CH848 SOSIP trimer
Method: single particle / : Manne K, Edwards RJ, Acharya P

EMDB-23094: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to one copy of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

EMDB-23095: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to two copies of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

EMDB-23097: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound domain-swapped antibody 2G12 from masked 3D refinement
Method: single particle / : Manne K, Henderson R, Acharya P

PDB-7l02: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to one copy of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

PDB-7l06: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound to two copies of domain-swapped antibody 2G12
Method: single particle / : Manne K, Henderson R, Acharya P

PDB-7l09: 
Cryo-EM structure of SARS-CoV-2 2P S ectodomain bound domain-swapped antibody 2G12 from masked 3D refinement
Method: single particle / : Manne K, Henderson R, Acharya P

EMDB-7490: 
1.71 A MicroED structure of proteinase K at 0.86 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7491: 
2.00 A MicroED structure of proteinase K at 2.6 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7492: 
2.20 A MicroED structure of proteinase K at 4.3 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7493: 
2.80 A MicroED structure of proteinase K at 6.0 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7494: 
3.20 A MicroED structure of proteinase K at 7.8 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7495: 
1.01 A MicroED structure of GSNQNNF at 0.27 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7496: 
1.01 A MicroED structure of GSNQNNF at 0.81 e- / A^2
Method: electron crystallography / : Hattne J, Shi D
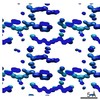
EMDB-7497: 
1.01 A MicroED structure of GSNQNNF at 1.3 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7498: 
1.15 A MicroED structure of GSNQNNF at 1.9 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7499: 
1.35 A MicroED structure of GSNQNNF at 2.4 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7500: 
1.37 A MicroED structure of GSNQNNF at 2.9 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7501: 
1.01 A MicroED structure of GSNQNNF at 0.17 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7502: 
1.01 A MicroED structure of GSNQNNF at 0.50 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7503: 
1.01 A MicroED structure of GSNQNNF at 0.82 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7504: 
1.02 A MicroED structure of GSNQNNF at 1.2 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7505: 
1.01 A MicroED structure of GSNQNNF at 1.5 e- / A^2
Method: electron crystallography / : Hattne J, Shi D
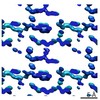
EMDB-7506: 
1.15 A MicroED structure of GSNQNNF at 1.8 e- / A^2
Method: electron crystallography / : Hattne J, Shi D
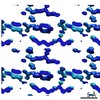
EMDB-7507: 
1.15 A MicroED structure of GSNQNNF at 2.1 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7508: 
1.16 A MicroED structure of GSNQNNF at 2.5 e- / A^2
Method: electron crystallography / : Hattne J, Shi D
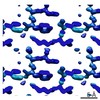
EMDB-7509: 
1.21 A MicroED structure of GSNQNNF at 2.8 e- / A^2
Method: electron crystallography / : Hattne J, Shi D
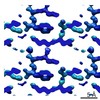
EMDB-7510: 
1.31 A MicroED structure of GSNQNNF at 3.1 e- / A^2
Method: electron crystallography / : Hattne J, Shi D
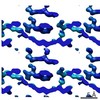
EMDB-7511: 
1.46 A MicroED structure of GSNQNNF at 3.4 e- / A^2
Method: electron crystallography / : Hattne J, Shi D
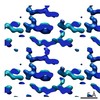
EMDB-7512: 
1.45 A MicroED structure of GSNQNNF at 3.8 e- / A^2
Method: electron crystallography / : Hattne J, Shi D

EMDB-7017: 
Segment from bank vole prion protein 168-176 QYNNQNNFV
Method: electron crystallography / : Glynn C, Rodriguez JA

EMDB-7287: 
MicroED structure of carbamazepine at 0.85 A resolution
Method: electron crystallography / : Gallagher-Jones M, Glynn C, Boyer DR, Martynowycz MW, Hernandez E, Miao J, Zee CT, NoviKova IV, Goldschmidt L, McFarlane HT, Helguera GF, Evans JE, Sawaya MR, Cascio D, Eisenberg D, Gonen T, Rodriguez JA
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
