-Search query
-Search result
Showing all 41 items for (author: huang & yx)

EMDB-39107: 
SARS-CoV-2 DMV nsp3-4 pore complex (full-pore)

EMDB-39109: 
SARS-CoV-2 DMV nsp3-4 pore complex (consensus-pore, C6 symmetry)

EMDB-39111: 
SARS-CoV-2 DMV nsp3-4 pore complex (extended-pore)

EMDB-39112: 
SARS-CoV-2 DMV nsp3-4 pore complex (consensus-pore, C3 symmetry)

EMDB-39113: 
SARS-CoV-2 DMV nsp3-4 pore complex (mini-pore)

EMDB-39159: 
SARS-CoV-2 DMV nsp3-4 pore complex (full-length-pore)

EMDB-33990: 
Cryo-EM structure of EBV gHgL-gp42 in complex with mAbs 3E8 and 5E3 (localized refinement)
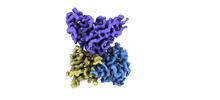
EMDB-33992: 
Cryo-EM structure of EBV gHgL-gp42 in complex with mAb 10E4 (localized refinement)
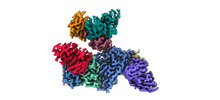
EMDB-33993: 
Cryo-EM density map of EBV gHgL-gp42 in complex with four mAbs 5E3, 3E8, 6H2 and 10E4
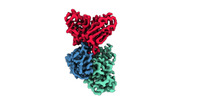
EMDB-33994: 
Cryo-EM structure of EBV gHgL-gp42 in complex with mAb 6H2 (localized refinement)
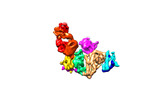
EMDB-33102: 
Cryo-EM structure of EBV glycoprotein complex gHgL-gp42 bound by a neutralizing antibody 6H2
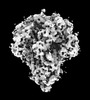
EMDB-32329: 
Cryo-EM map of PEDV (Pintung 52) S protein with all three protomers in the D0-down conformation determined in situ on intact viral particles.

EMDB-32332: 
Subtomogram averaging of PEDV (Pintung 52) S protein with all three protomers in the D0-down conformation determined in situ on intact viral particles.

EMDB-32333: 
Subtomogram averaging of PEDV (Pintung 52) S protein with one protomer in the D0-up conformation and two protomers in the D0-down conformation, determined in situ on intact viral particles

EMDB-32337: 
Subtomogram averaging of PEDV (Pintung 52) S protein with two protomers in the D0-up conformation and one protomer in the D0-down conformation, determined in situ on intact viral particles.

EMDB-32338: 
Cryo-EM map of PEDV S protein with one protomer in the D0-up conformation while the other two in the D0-down conformation
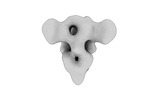
EMDB-32339: 
Subtomogram averaging of PEDV (Pintung 52) S protein with all three protomers in the D0-up conformation determined in situ on intact viral particles.

EMDB-32340: 
Subtomogram averaging of PEDV (Pintung 52) S protein in the postfusion form determined in situ on intact viral particles.

EMDB-33646: 
Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein with three D0-up

EMDB-33647: 
Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein one D0-down and two D0-up

EMDB-33648: 
Symmetry-expanded and locally refined protomer structure of IPEC-J2 cell-derived PEDV PT52 S with a CTD-close conformation
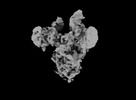
EMDB-33649: 
Symmetry-expanded and locally refined protomer structure of IPEC-J2 cell-derived PEDV PT52 S with a CTD-open conformation

EMDB-33700: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein with three D0-down
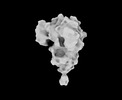
EMDB-33701: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein one D0-up and two D0-down

EMDB-33702: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein with three D0-up
EMDB-33703: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I with three D0-down

EMDB-33704: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I one D0-up and two D0-down
EMDB-33705: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I one D0-down and two D0-up
EMDB-33706: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I with three D0-up

EMDB-32557: 
SARS-CoV-2 Omicron S-open

EMDB-32170: 
SARS-CoV-2 Beta variant spike protein in transition state

EMDB-31146: 
Coupling of N7-methyltransferase and 3'-5' exoribonuclease with SARS-CoV-2 polymerase reveals mechanisms for capping and proofreading

EMDB-31138: 
Co-transcriptional capping machineries in SARS-CoV-2 RTC: Coupling of N7-methyltransferase and 3'-5' exoribonuclease with polymerase reveals mechanisms for capping and proofreading

EMDB-21249: 
Cryo-EM structure of an activated VIP1 receptor-G protein complex

EMDB-20417: 
Cryo-EM reconstruction of the mature chimeric BinJV/ZIKV-prME virion

EMDB-20438: 
Cryo-EM reconstruction of the mature chimeric BinJV/ZIKV-prME virion in complex with Fab C8

EMDB-20439: 
Cryo-EM reconstruction of the immature chimeric BinJV/ZIKV-prME virion

EMDB-20416: 
Cryo-EM structure of the full-length Bacillus subtilis glyQS T-box riboswitch in complex with tRNA-Gly

EMDB-9694: 
The complete structure of yeast COMPASS

EMDB-2870: 
Cryo-EM structure of the multimeric complex between Dark and Dronc-CARD.

EMDB-2871: 
Cryo-EM structure of the Dark apoptosome.
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
