-Search query
-Search result
Showing 1 - 50 of 55 items for (author: eiler & dr)

EMDB-29721: 
40S ribosomal subunit of the 80S Giardia intestinalis assemblage A ribosome with Emetine bound in V1 conformation
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB-29897: 
Focused refinement of the 40S subunit head of the 80S Giardia lamblia ribosome at 2.94 angstroms resolution.
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB-29906: 
80S Giardia lamblia ribosome at 2.66 angstroms resolution with Emetine in the the V1 conformation
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB-29407: 
60S subunit of the Giardia lamblia 80S ribosome
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB-29730: 
40S ribosomal subunit of the 80S Giardia intestinalis assemblage A ribosome with Emetine bound in V2 conformation with mRNA and three tRNAs.
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB-29918: 
80S Giardia lamblia ribosome at 2.67 angstroms resolution with Emetine in the the V2 conformation
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB-29495: 
40S subunit of the Giardia lamblia 80S ribosome
Method: single particle / : Eiler DR, Wimberly BT, Bilodeau DY, Rissland OS, Kieft JS

EMDB-26735: 
Hantavirus ANDV Gn(H) protein in complex with 2 Fabs ANDV-5 and ANDV-34
Method: single particle / : Binshtein E, Crowe JE

EMDB-26736: 
Hantavirus MAPV Gn(H)/Gc protein in complex with 2 Fabs SNV-24 and SNV-53
Method: single particle / : Binshtein E, Crowe JE

EMDB-27318: 
CryoEM structure of Hantavirus ANDV Gn(H) protein complex with 2Fabs ANDV-5 and ANDV-34
Method: single particle / : Binshtein E, Crowe JE

PDB-8dbz: 
CryoEM structure of Hantavirus ANDV Gn(H) protein complex with 2Fabs ANDV-5 and ANDV-34
Method: single particle / : Binshtein E, Crowe JE

EMDB-16686: 
CupE pilus (CupE1 111-113 AGA mutant)
Method: helical / : Boehning J, Bharat TAM

EMDB-16683: 
Cryo-EM structure of the CupE pilus from Pseudomonas aeruginosa
Method: helical / : Boehning J, Bharat TAM

PDB-8cio: 
Cryo-EM structure of the CupE pilus from Pseudomonas aeruginosa
Method: helical / : Boehning J, Bharat TAM

EMDB-21965: 
Negative stain EM map of SARS CoV-2 spike protein (trimer)
Method: single particle / : Binshtein E

EMDB-21974: 
Negative stain EM map of SARS-CoV-2 spike protein (trimer) with Fab COV2-2165
Method: single particle / : Binshtein E

EMDB-21975: 
Negative stain EM map of SARS-CoV-2 spike protein (trimer) with Fab COV2-2196
Method: single particle / : Binshtein E

EMDB-21976: 
Negative stain EM map of SARS-CoV-2 spike protein (trimer) with Fab COV2-2130
Method: single particle / : Binshtein E

EMDB-21977: 
Negative stain EM map of SARS-CoV-2 spike protein (trimer) with Fab COV2-2130 and Fab COV2-2196
Method: single particle / : Binshtein E
EMDB-7908: 
VIC170 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7909: 
VIC169 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
EMDB-7910: 
VIC167 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
EMDB-7911: 
VIC166 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
EMDB-7912: 
VIC165 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7914: 
VIC1164 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
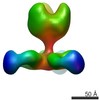
EMDB-7915: 
VIC163 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7926: 
VIC82 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
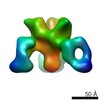
EMDB-7927: 
VIC43 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7928: 
VIC32 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7929: 
VIC26 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7930: 
VIC16 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7931: 
VIC15 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7932: 
VIC12 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
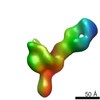
EMDB-7933: 
VIC11 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7922: 
VIC133 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7923: 
VIC101 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7924: 
VIC94 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7925: 
VIC91 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7934: 
VIC130 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
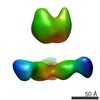
EMDB-7916: 
VIC161 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7917: 
VIC160 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
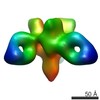
EMDB-7918: 
VIC158 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
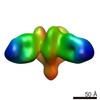
EMDB-7919: 
VIC157 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-7920: 
VIC142 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB
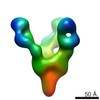
EMDB-7921: 
VIC139 Fab in complex with Ebola virus GP
Method: single particle / : Turner H, Murin CD, Pallesen J, Ward AB

EMDB-2448: 
Cryo-EM structure of T. thermophilus 30S Translation Initiation complex
Method: single particle / : Simonetti A, Marzi S, Billas IML, Tsai A, Fabbretti A, Myasnikov A, Roblin P, Vaiana AC, Hazemann I, Eiler D, Steitz TA, Puglisi JD, Gualerzi GO, Klaholz BP

EMDB-2523: 
Structure of the mammalian oligosaccharyl-transferase complex in the native ER protein translocon
Method: single particle / : Pfeffer S, Dudek J, Gogala M, Schorr S, Linxweiler J, Lang S, Becker T, Beckmann R, Zimmermann F

EMDB-2514: 
ER membrane-associated ribosome from HeLa cells after treatment with non-silencing control siRNA
Method: subtomogram averaging / : Pfeffer S, Dudek J, Gogala M, Schorr S, Linxweiler J, Lang S, Becker T, Beckmann R, Zimmermann R, Foerster F

EMDB-2515: 
ER membrane-associated ribosome from HeLa cells after treatment with TRAP-beta siRNA
Method: subtomogram averaging / : Pfeffer S, Dudek J, Gogala M, Schorr S, Linxweiler J, Lang S, Becker T, Beckmann R, Zimmermann R, Foerster F

EMDB-2516: 
ER membrane-associated ribosome from HeLa cells after treatment with Ribophorin I siRNA
Method: subtomogram averaging / : Pfeffer S, Dudek J, Gogala M, Schorr S, Linxweiler J, Lang S, Becker T, Beckmann R, Zimmermann R, Foerster F
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
