[English] 日本語
 Yorodumi
Yorodumi- EMDB-2523: Structure of the mammalian oligosaccharyl-transferase complex in ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2523 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the mammalian oligosaccharyl-transferase complex in the native ER protein translocon | |||||||||
 Map data Map data | Reconstruction was masked | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Co-translational protein translocation / N-Glycosylation | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 9.3 Å | |||||||||
 Authors Authors | Pfeffer S / Dudek J / Gogala M / Schorr S / Linxweiler J / Lang S / Becker T / Beckmann R / Zimmermann F | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2014 Journal: Nat Commun / Year: 2014Title: Structure of the mammalian oligosaccharyl-transferase complex in the native ER protein translocon. Authors: Stefan Pfeffer / Johanna Dudek / Marko Gogala / Stefan Schorr / Johannes Linxweiler / Sven Lang / Thomas Becker / Roland Beckmann / Richard Zimmermann / Friedrich Förster /  Abstract: In mammalian cells, proteins are typically translocated across the endoplasmic reticulum (ER) membrane in a co-translational mode by the ER protein translocon, comprising the protein-conducting ...In mammalian cells, proteins are typically translocated across the endoplasmic reticulum (ER) membrane in a co-translational mode by the ER protein translocon, comprising the protein-conducting channel Sec61 and additional complexes involved in nascent chain processing and translocation. As an integral component of the translocon, the oligosaccharyl-transferase complex (OST) catalyses co-translational N-glycosylation, one of the most common protein modifications in eukaryotic cells. Here we use cryoelectron tomography, cryoelectron microscopy single-particle analysis and small interfering RNA-mediated gene silencing to determine the overall structure, oligomeric state and position of OST in the native ER protein translocon of mammalian cells in unprecedented detail. The observed positioning of OST in close proximity to Sec61 provides a basis for understanding how protein translocation into the ER and glycosylation of nascent proteins are structurally coupled. The overall spatial organization of the native translocon, as determined here, serves as a reliable framework for further hypothesis-driven studies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2523.map.gz emd_2523.map.gz | 10.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2523-v30.xml emd-2523-v30.xml emd-2523.xml emd-2523.xml | 10.7 KB 10.7 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2423.jpg EMD-2423.jpg | 71 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2523 http://ftp.pdbj.org/pub/emdb/structures/EMD-2523 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2523 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2523 | HTTPS FTP |
-Related structure data
| Related structure data |  2514C  2515C  2516C  2517C  2518C  2519C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_2523.map.gz / Format: CCP4 / Size: 804.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2523.map.gz / Format: CCP4 / Size: 804.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction was masked | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.2374 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
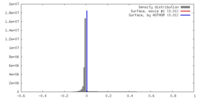
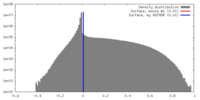
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Canis familiaris Sec61, OST and TRAP bound to a translocating whe...
| Entire | Name: Canis familiaris Sec61, OST and TRAP bound to a translocating wheat germ 80S ribosome |
|---|---|
| Components |
|
-Supramolecule #1000: Canis familiaris Sec61, OST and TRAP bound to a translocating whe...
| Supramolecule | Name: Canis familiaris Sec61, OST and TRAP bound to a translocating wheat germ 80S ribosome type: sample / ID: 1000 Oligomeric state: One 80S ribosome binds one Sec61, one OST and one TRAP Number unique components: 4 |
|---|
-Supramolecule #1: Triticum aestivum 80S ribosome
| Supramolecule | Name: Triticum aestivum 80S ribosome / type: complex / ID: 1 / Name.synonym: Wheat germ 80S ribosome / Details: Data-subset resulted from computational sorting / Recombinant expression: No / Ribosome-details: ribosome-eukaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Sec61
| Macromolecule | Name: Sec61 / type: protein_or_peptide / ID: 1 / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #2: Oligosaccharyltransferase
| Macromolecule | Name: Oligosaccharyltransferase / type: protein_or_peptide / ID: 2 / Name.synonym: OST / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #3: Translocon-Associated Protein omplex
| Macromolecule | Name: Translocon-Associated Protein omplex / type: protein_or_peptide / ID: 3 / Name.synonym: TRAP / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.6 Details: 30 mM Hepes/KOH 7.6, 10 mM Mg(OAc)2, 180 mM KOAC/HAc pH 7.6, 0.3 % Digitonin, 1 mM DTT |
|---|---|
| Grid | Details: Quantifoil grids pre-coated with 2 nm carbon on top |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Instrument: FEI VITROBOT MARK IV / Method: Blot for 3 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Date | Jul 17, 2011 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Number real images: 6119 / Average electron dose: 25 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 148721 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.3 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | The particles were selected using SIGNATURE, classified using MAPPOS and processed with SPIDER |
|---|---|
| CTF correction | Details: On 3D-volume (SPIDER) |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 9.3 Å / Resolution method: OTHER / Software - Name: SPIDER, Signature, MAPPOS Details: This datasubset resulted from computational sorting. Number images used: 15705 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Software | Name: Chimera, Coot, MDFF |
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) X (Row.)
X (Row.) Y (Col.)
Y (Col.)