-検索条件
-検索結果
検索 (著者・登録者: efremov & r)の結果114件中、1から50件目までを表示しています

EMDB-17112: 
Structure of apo form of human gamma-secretase PSEN1 APH-1B isoform reconstituted into lipid nanodisc

EMDB-17113: 
Structure of human gamma-secretase PSEN1 APH-1B isoform reconstituted into lipid nanodisc in complex with Ab46

PDB-8oqy: 
Structure of apo form of human gamma-secretase PSEN1 APH-1B isoform reconstituted into lipid nanodisc

PDB-8oqz: 
Structure of human gamma-secretase PSEN1 APH-1B isoform reconstituted into lipid nanodisc in complex with Ab46

EMDB-16157: 
Map of GroEL:GroES-(ADP) complex plunge frozen 200 ms after reaction initiation with ATP

EMDB-40676: 
Human TRP channel TRPV6 in cNW30 nanodiscs inhibited by tetrahydrocannabivarin (THCV)

PDB-8sp8: 
Human TRP channel TRPV6 in cNW30 nanodiscs inhibited by tetrahydrocannabivarin (THCV)

EMDB-16091: 
Cryo-EM structure of beta-galactosidase at 3.3 A resolution plunged 5 ms after mixing with apoferritin

EMDB-16092: 
Cryo-EM structure of beta-galactosidase at 2.9 A resolution plunged 205 ms after mixing with apoferritin

EMDB-16093: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.1 A plunged 5ms after mixing with b-galactosidase

EMDB-16094: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.7 A plunged 35ms after mixing with b-galactosidase

EMDB-16095: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.2 A plunged 205ms after mixing with b-galactosidase

EMDB-16097: 
Cryo-EM structure of beta-galactosidase at 3.2 A resolution plunged 35 ms after mixing with apoferritin
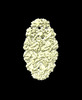
EMDB-16099: 
GroEL:GroES-ATP complex under continuous turnover condirions

EMDB-16100: 
Structure of GroEL-ATP complex plunge frozen 200 ms after reaction initiation

EMDB-16102: 
Structure of GroEL-nucleotide complex in ADP-like conformation plunged 13 ms after mixing with ATP

EMDB-16106: 
Structure of the GroEL-ATP complex plunge-frozen 50 ms after mixing with ATP

EMDB-16107: 
Structure of the GroEL(ATP7/ADP7) complex plunged 13 ms after mixing with ATP

EMDB-16108: 
Structure of GroEL-nucleotide complex in ADP-like conformation plunged 50 ms after mixing with ATP

EMDB-16109: 
Structure of the GroEL(ATP7/ADP7) complex plunged 50 ms after mixing with ATP

EMDB-16115: 
Structure of the GroEL-ATP complex plunge-frozen 13 ms after mixing with ATP

EMDB-16116: 
Structure of GroEL:GroES-ATP complex plunge frozen 200 ms after reaction initiation

EMDB-16117: 
Structure of GroEL:GroES-ATP complex under continuous turnover conditions

EMDB-16118: 
Structure of GroEL-ATP complex under continuous turnover conditions

EMDB-16119: 
Structure of GroEL:GroES complex exhibiting ADP-conformation in trans ring obtained under the continuous turnover conditions

EMDB-16125: 
Structure of GroEL:GroES-ATP complex plunge frozen 200 ms after reaction initiation

EMDB-16154: 
Map of the GroEL-ES-ATP complex plunge-frozen 50 ms after mixing with ATP

PDB-8bk7: 
Cryo-EM structure of beta-galactosidase at 3.3 A resolution plunged 5 ms after mixing with apoferritin

PDB-8bk8: 
Cryo-EM structure of beta-galactosidase at 2.9 A resolution plunged 205 ms after mixing with apoferritin

PDB-8bk9: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.1 A plunged 5ms after mixing with b-galactosidase

PDB-8bka: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.7 A plunged 35ms after mixing with b-galactosidase

PDB-8bkb: 
Cryo-EM structure of mouse heavy-chain apoferritin at 2.2 A plunged 205ms after mixing with b-galactosidase

PDB-8bkg: 
Cryo-EM structure of beta-galactosidase at 3.2 A resolution plunged 35 ms after mixing with apoferritin

PDB-8bkz: 
GroEL:GroES-ATP complex under continuous turnover conditions

PDB-8bl2: 
Structure of GroEL-ATP complex plunge frozen 200 ms after reaction initiation

PDB-8bl7: 
Structure of GroEL-nucleotide complex in ADP-like conformation plunged 13 ms after mixing with ATP

PDB-8blc: 
Structure of the GroEL-ATP complex plunge-frozen 50 ms after mixing with ATP

PDB-8bld: 
Structure of the GroEL(ATP7/ADP7) complex plunged 13 ms after mixing with ATP

PDB-8ble: 
Structure of GroEL-nucleotide complex in ADP-like conformation plunged 50 ms after mixing with ATP

PDB-8blf: 
Structure of the GroEL(ATP7/ADP7) complex plunged 50 ms after mixing with ATP

PDB-8bly: 
Structure of the GroEL-ATP complex plunge-frozen 13 ms after mixing with ATP

PDB-8bm0: 
Structure of GroEL:GroES-ATP complex plunge frozen 200 ms after reaction initiation

PDB-8bm1: 
Structure of GroEL:GroES-ATP complex under continuous turnover conditions

PDB-8bmd: 
Structure of GroEL-ATP complex under continuous turnover conditions

PDB-8bmo: 
Structure of GroEL:GroES complex exhibiting ADP-conformation in trans ring obtained under the continuous turnover conditions

PDB-8bmt: 
Structure of GroEL:GroES-ATP complex plunge frozen 200 ms after reaction initiation
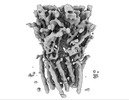
EMDB-16308: 
Alvinella pompejana nicotinic acetylcholine receptor Alpo4 in apo state (dataset 1)
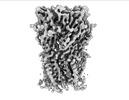
EMDB-16314: 
Alvinella pompejana nicotinic acetylcholine receptor Alpo in apo state (dataset 2)
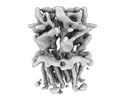
EMDB-16315: 
Alvinella pompejana nicotinic acetylcholine receptor Alpo4 in apo state (Alpo4_LMNG_Serotonin dataset 4)
EMDB-16316: 
Alvinella pompejana nicotinic acetylcholine receptor Alpo4 in apo state (Alpo4_comb dataset 3)
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
