-Search query
-Search result
Showing all 41 items for (author: chen & zn)

EMDB-66696: 
Human KCNQ2-CaM in complex with QO-58 and PIP2
Method: single particle / : Zhao YW, Yang ZN, Guo JT, Du XN

EMDB-66788: 
Human KCNQ2-CaM in complex with QO-83 and PIP2
Method: single particle / : Zhao YW, Yang ZN, Du XN, Guo JT

EMDB-62892: 
Human KCNQ2-CaM in complex with QO-58
Method: single particle / : Zhao YW, Yang ZN, Du XN, Guo JT

EMDB-64131: 
Structure of SARS-CoV-2 spike-CD147 complex at 3.75 Angstroms resolution
Method: single particle / : Zhang SJ, Yang ZW, Lin P, Bian HJ, Zhu P, Zhang L, Chen ZN

EMDB-38522: 
Human KCNQ2-CaM in complex with QO-83
Method: single particle / : Zhao YW, Yang ZN, Guo JT, Du XN

EMDB-44438: 
Cryo-EM structure of Thermococcus kodakarensis FttA-dependent transcription pre-termination complex containing 44 nt RNA
Method: single particle / : You L, Ebright RH

EMDB-44439: 
Cryo-EM structure of Thermococcus kodakarensis FttA-dependent transcription pre-termination complex containing 52 nt RNA
Method: single particle / : You L, Ebright RH

EMDB-44454: 
Cryo-EM structure of Thermococcus kodakarensis FttA-dependent transcription pre-termination complex containing 44 nt RNA
Method: single particle / : You L, Ebright RH

EMDB-44455: 
Cryo-EM structure of Thermococcus kodakarensis FttA-dependent transcription pre-termination complex containing 52 nt RNA
Method: single particle / : You L, Ebright RH

EMDB-44649: 
Cryo-EM structure of Thermococcus kodakarensis FttA-dependent transcription pre-termination complex containing 44 nt RNA (local refinement map)
Method: single particle / : You L, Ebright RH

EMDB-44650: 
Cryo-EM structure of Thermococcus kodakarensis FttA-dependent transcription pre-termination complex containing 52 nt RNA (local refinement map)
Method: single particle / : You L, Ebright RH

PDB-9bct: 
Cryo-EM structure of Thermococcus kodakarensis FttA-dependent transcription pre-termination complex containing 44 nt RNA
Method: single particle / : You L, Ebright RH

PDB-9bcu: 
Cryo-EM structure of Thermococcus kodakarensis FttA-dependent transcription pre-termination complex containing 52 nt RNA
Method: single particle / : You L, Ebright RH

EMDB-37006: 
Cryo-EM structure of SARS-CoV-2 Delta RBD in complex with golden hamster ACE2 (local refinement)
Method: single particle / : Niu S, Zhao ZN, Chai Y, Gao GF

EMDB-37090: 
Cryo-EM structure of SARS-CoV-2 BA.3 RBD in complex with golden hamster ACE2 (local refinement)
Method: single particle / : Niu S, Zhao ZN, Chai Y, Gao GF

EMDB-15165: 
Structure of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of Chaetomium thermophilum INO80
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

PDB-8a5d: 
Structure of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of Chaetomium thermophilum INO80
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15163: 
Cryo-EM reconstruction of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of INO80
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15164: 
Human Ino80 A-module + YY1
Method: single particle / : Kunert F, Metzner FJ, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15177: 
Cryo-EM reconstruction of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of S. cerevisiae INO80
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15179: 
Cryo-EM reconstruction of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of S. cerevisiae INO80
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15180: 
Structure of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of Chaetomium thermophilum INO80 on curved DNA
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15184: 
Structure of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of Chaetomium thermophilum INO80 on straight DNA
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15186: 
Cryo-EM reconstruction of DNA bound Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of S. cerevisiae INO80
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15187: 
A-module of Chaetomium thermophilum INO80 bound to curved DNA
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15188: 
Nucleosome bound Chaetomium thermophilum INO80 (ADP-BeF3 state) core complex with Arp5 grappler in parallel conformation
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15211: 
Ino80 core complex with A-module on nucleosome
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15647: 
Nucleosome-bound Ino80 ATPase
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-15688: 
CryoEM structure of INO80 core nucleosome complex in closed grappler conformation (ADP-BeFx state)
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

PDB-8a5a: 
Structure of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of INO80
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

PDB-8a5o: 
Structure of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of S. cerevisiae INO80
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

PDB-8a5p: 
Structure of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of Chaetomium thermophilum INO80 on curved DNA
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

PDB-8a5q: 
Structure of Arp4-Ies4-N-actin-Arp8-Ino80HSA subcomplex (A-module) of Chaetomium thermophilum INO80 on straight DNA
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

PDB-8atf: 
Nucleosome-bound Ino80 ATPase
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

PDB-8av6: 
CryoEM structure of INO80 core nucleosome complex in closed grappler conformation
Method: single particle / : Kunert F, Metzner FJ, Eustermann S, Jung J, Woike S, Schall K, Kostrewa D, Hopfner KP

EMDB-33697: 
Cryo-EM structure of SARS-CoV-2 Omicron spike protein (S-6P-RRAR) in complex with human ACE2 ectodomain (one-RBD-up state)
Method: single particle / : Gao GF, Qi JX, Liu S, Zhao ZN

EMDB-33698: 
Cryo-EM structure of hACE2-bound SARS-CoV-2 Omicron spike protein with L371S, P373S and F375S mutations (S-6P-RRAR)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-33690: 
Cryo-EM structure of apo SARS-CoV-2 Omicron spike protein (S-2P-GSAS)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-33699: 
Cryo-EM structure of hACE2-bound SARS-CoV-2 Omicron spike protein with L371S, P373S and F375S mutations (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao GF

EMDB-33709: 
Cryo-EM structure of S309-RBD-RBD-S309 in the S309-bound Omicron spike protein (local refinement)
Method: single particle / : Zhao ZN, Xie YF, Qi JX, Gao F
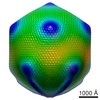
EMDB-5039: 
Cryo-EM reconstruction of the giant Mimivirus using C5 symmetry
Method: single particle / : Xiao C, Kuznetsov YG, Sun S, Hafenstein SL, Kostyuchenko VA, Chipman PR, Suzan-Monti M, Raoult D, McPherson A, Rossmann MG
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
