-Search query
-Search result
Showing all 33 items for (author: cassidy & ck)

EMDB-16489: 
In situ structure of the Nitrosopumilus maritimus S-layer - Six-fold symmetry (C6)

EMDB-16492: 
In situ structure of the Nitrosopumilus maritimus S-layer - Composite map between C2 and C6

EMDB-16482: 
In vitro structure of the Nitrosopumilus maritimus S-layer - Six-fold symmetry (C6)

EMDB-16483: 
In vitro structure of the Nitrosopumilus maritimus S-layer - Two-fold symmetry (C2)

EMDB-16484: 
In vitro structure of the Nitrosopumilus maritimus S-layer - Composite map between two and six-fold symmetrised

EMDB-16486: 
In vitro Nitrosopumilus maritimus S-layer with NH4Cl

EMDB-16487: 
In situ structure of the Nitrosopumilus maritimus S-layer - Two-fold symmetry (C2)

EMDB-15648: 
Structure of the giant inhibitor of apoptosis, BIRC6 (multibody map 1)

EMDB-15650: 
Structure of the giant inhibitor of apoptosis, BIRC6 (multibody map 2)

EMDB-15651: 
Structure of the giant inhibitor of apoptosis, BIRC6 (multibody map 3)

EMDB-15652: 
Structure of the giant inhibitor of apoptosis, BIRC6 (homogenous refinement)

EMDB-15653: 
Structure of the giant inhibitor of apoptosis, BIRC6 (composite map)
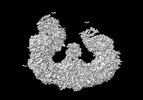
EMDB-15654: 
Structure of the giant inhibitor of apoptosis, BIRC6 bound to the regulator SMAC

EMDB-15641: 
Structure of the Native Chemotaxis Core Signalling Complex from E-gene lysed E. coli cells.

EMDB-15642: 
Native Chemotaxis Core Signalling Complex from E. coli, with focused alignment on the baseplate (CheA-CheW)
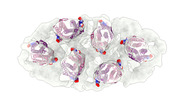
EMDB-15643: 
Native Chemotaxis Core Signalling Complex from E. coli, Focused alignment on ligand binding domain

EMDB-15669: 
Jumbo Phage phi-kp24 tail outer sheath

EMDB-14356: 
Jumbo Phage phi-Kp24 full capsid

EMDB-14357: 
Jumbo Phage phi-Kp24 extended tail
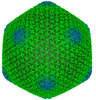

EMDB-13862: 
Jumbo Phage phi-Kp24 empty capsid

EMDB-14399: 
pMMO structure from native membranes by cryoET and STA

EMDB-14530: 
pMMO three trimer interaction map from native membrane

EMDB-13364: 
UVC treated Human apoferritin

EMDB-13402: 
Cryo-EM map of UVC-treated ICP1 Bacteriophage capsid

EMDB-13403: 
Cryo-EM map of WT ICP1 bacteriophage capsid

EMDB-10050: 
Structure of the E. coli Chemotaxis Core Signaling Unit

EMDB-10160: 
In Situ Core-Signalling Unit of E. coli Chemoreceptor Array

EMDB-4991: 
Escherichia coli chemotaxis signaling arrays at low kinase activity with serine receptor mutant Tsr_EEEE

EMDB-4992: 
Escherichia coli chemotaxis signaling arrays at high kinase activity with serine receptor mutant Tsr_QQQQ

EMDB-4993: 
Escherichia coli chemotaxis signaling arrays with wild-type serine receptor Tsr_QEQE

EMDB-6319: 
Structure of bacterial chemotaxis signaling CheA2-trimer core complex by cryo-electron tomography and subvolume averaging

EMDB-6320: 
Structure of bacterial chemotaxis signaling CheA2-hexamer core complex by cryo-electron tomography and subvolume averaging

EMDB-3234: 
Representative tomogram as used in: Structure of bacterial chemotaxis signaling CheA2-trimer core complex by cryo-electron tomography and subvolume averaging
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
