-検索条件
-検索結果
検索 (著者・登録者: bohn & s)の結果121件中、1から50件目までを表示しています

EMDB-18779: 
Structure of the non-mitochondrial citrate synthase from Ananas comosus

PDB-8qzp: 
Structure of the non-mitochondrial citrate synthase from Ananas comosus

EMDB-17418: 
CryoEM structure of a C7-symmetrical GroEL7-GroES7 cage in presence of ADP-BeFx

EMDB-17420: 
CryoEM structure of a GroEL7-GroES7 cage with encapsulated disordered substrate MetK in the presence of ADP-BeFx

EMDB-17421: 
CryoEM structure of a GroEL7-GroES7 cage with encapsulated ordered substrate MetK in the presence of ADP-BeFx

EMDB-17422: 
Density for MetK encapsulated in the GroEL7-GroES7 cage

EMDB-17423: 
Symmetry-averaged GroEL7-GroES7 chamber with encapsulated disordered substrate MetK obtained by in vitro cryo electron tomography
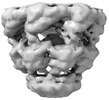
EMDB-17424: 
Symmetry-averaged GroEL7-GroES7 chamber with encapsulated ordered substrate MetK obtained by in vitro cryo electron tomography

EMDB-17425: 
Structure average of GroEL14 complexes found in the cytosol of Escherichia coli overexpressing GroEL obtained by cryo electron tomography

EMDB-17426: 
In situ structure average of GroEL14-GroES14 complexes in Escherichia coli cytosol obtained by cryo electron tomography

EMDB-17534: 
Cryo-EM structure of a D7-symmetrical GroEL14-GroES14 complex in presence of ADP-BeFx

EMDB-17535: 
Cryo-EM structure of a C7-symmetrical GroEL14-GroES7 complex in presence of ADP-BeFx

EMDB-17559: 
In situ structure average of GroEL7-GroES7 chamber with no or disordered substrate in Escherichia coli cytosol obtained by cryo electron tomography

EMDB-17560: 
In situ structure average of GroEL7-GroES7 chamber with encapsulated, ordered substrate in Escherichia coli cytosol obtained by cryo electron tomography
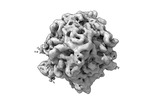
EMDB-17561: 
Cryo-ET subtomogram of 70S ribosomes in Escherichia coli cells at 37 and 46 degrees centigrade and in Escherichia coli cells overexpressing GroELS and MetK

EMDB-17562: 
Cryo-ET subtomogram of 70S ribosomes in Escherichia coli cells overexpressing GroEL

EMDB-17563: 
CryoEM structure of a GroEL14-GroES7 cage with encapsulated ordered substrate MetK in the presence of ADP-BeFx

EMDB-17564: 
CryoEM structure of a GroEL14-GroES7 cage with encapsulated disordered substrate MetK in the presence of ADP-BeFx

EMDB-17565: 
CryoEM structure of a GroEL14-GroES14 cage with two encapsulated disordered MetK substrates in the presence of ADP-BeFx

EMDB-17566: 
CryoEM structure of a (GroEL)14-(GroES)14 complex with encapsulated ordered MetK substrate in one chamber and no or disordered MetK substrate in the other chamber in the presence of ADP-BeFx
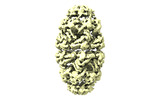
EMDB-17567: 
Conformer 1 of the (GroEL)14(GroES)14 complex with two encapsulated, ordered and near-native MetK substrate molecules in the presence of ADP-BeFx

EMDB-17568: 
Conformer 2 of the (GroEL)14(GroES)14 complex with two encapsulated, near-native and ordered MetK substrates in the presence of ADP-BeFx
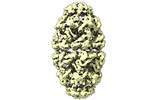
EMDB-17569: 
Conformer 3 of the (GroEL)14(GroES)14 complex with two encapsulated, near-native and ordered MetK substrates in the presence of ADP-BeFx

EMDB-17570: 
Conformer 4 of the (GroEL)14(GroES)14 complex with two encapsulated, near-native and ordered MetK substrates in the presence of ADP-BeFx

EMDB-17571: 
Conformer 5 of the (GroEL)14(GroES)14 complex with two encapsulated, near-native and ordered MetK substrates in the presence of ADP-BeFx

EMDB-17572: 
Conformer 6 of the (GroEL)14(GroES)14 complex with two encapsulated, near-native and ordered MetK substrates in the presence of ADP-BeFx

EMDB-17573: 
Conformer 7 of the (GroEL)14(GroES)14 complex with two encapsulated, near-native and ordered MetK substrates in the presence of ADP-BeFx

EMDB-18735: 
CryoEM structure of a GroEL14-GroES7 complex in presence of ADP-BeFx with wide GroEL7 trans ring conformation

EMDB-18736: 
CryoEM structure of a GroEL14-GroES7 complex in presence of ADP-BeFx with narrow GroEL7 trans ring conformation

EMDB-18737: 
In situ structure average of GroEL14-GroES7 complexes with wide GroEL7 trans ring conformation in Escherichia coli cytosol obtained by cryo electron tomography

EMDB-18738: 
In situ structure average of GroEL14-GroES7 complexes with narrow GroEL7 trans ring conformation in Escherichia coli cytosol obtained by cryo electron tomography

PDB-8p4m: 
CryoEM structure of a C7-symmetrical GroEL7-GroES7 cage in presence of ADP-BeFx

PDB-8p4n: 
CryoEM structure of a GroEL7-GroES7 cage with encapsulated disordered substrate MetK in the presence of ADP-BeFx

PDB-8p4o: 
CryoEM structure of a GroEL7-GroES7 cage with encapsulated ordered substrate MetK in the presence of ADP-BeFx

PDB-8p4p: 
Structure average of GroEL14 complexes found in the cytosol of Escherichia coli overexpressing GroEL obtained by cryo electron tomography

PDB-8p4r: 
In situ structure average of GroEL14-GroES14 complexes in Escherichia coli cytosol obtained by cryo electron tomography

PDB-8qxs: 
CryoEM structure of a GroEL14-GroES7 complex in presence of ADP-BeFx with wide GroEL7 trans ring conformation

PDB-8qxt: 
CryoEM structure of a GroEL14-GroES7 complex in presence of ADP-BeFx with narrow GroEL7 trans ring conformation

PDB-8qxu: 
In situ structure average of GroEL14-GroES7 complexes with wide GroEL7 trans ring conformation in Escherichia coli cytosol obtained by cryo electron tomography

PDB-8qxv: 
In situ structure average of GroEL14-GroES7 complexes with narrow GroEL7 trans ring conformation in Escherichia coli cytosol obtained by cryo electron tomography

EMDB-17924: 
Cryo-EM structure of human Elp123 in complex with tRNA, acetyl-CoA, 5'-deoxyadenosine and methionine

EMDB-17925: 
Cryo-EM structure of human Elp123 in complex with 5'-deoxyadenosine and methionine

EMDB-17926: 
Cryo-EM structure of human Elp123 in complex with tRNA, S-ethyl-CoA, 5'-deoxyadenosine and methionine

EMDB-17927: 
Cryo-EM structure of human Elp123 in complex with tRNA, desulpho-CoA, 5'-deoxyadenosine and methionine

EMDB-18892: 
Lipid droplet-Vacuole contacts in Ldo16 overexpression yeast strain.

EMDB-18893: 
Lipid droplet-vacuole and Nucleus-vacuole contacts in WT yeast cell starved for 4 hours

EMDB-18894: 
Lipid droplet lipophagy in 4-hour starved WT yeast cell.

EMDB-18895: 
Multiple vacuole-lipid droplet-nucleus contacts in 4-hour starved WT yeast cell.

EMDB-18896: 
Lipophagy in 5-day starved WT yeast cell.

EMDB-18897: 
Lipid droplets in proximity to a vacuole in dLdo strain cell after 5-day starvation.
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
