-Search query
-Search result
Showing 1 - 50 of 117 items for (author: baker & rw)

EMDB-71909: 
Structure of AP-2 bound to the dileucine motif of CCDC32; combined map
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-71914: 
Composite structure of AP-2 bound to the dileucine motif and WxxPhi motif of CCDC32
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-71906: 
Structure of AP-2 bound to the dileucine motif of CCDC32; consensus refinement
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-71911: 
Structure of AP-2 bound to the dileucine motif and WxxPhi motif of CCDC32; focused refinement 1
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-71912: 
Structure of AP-2 bound to the dileucine motif and WxxPhi motif of CCDC32; focused refinement 2
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-71913: 
Structure of AP-2 bound to the dileucine motif and WxxPhi motif of CCDC32; focused refinement 3
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-71905: 
Closed AP-2 clathrin adaptor complex in solution
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-71907: 
Structure of AP-2 bound to the dileucine motif of CCDC32; focused refinement
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-71910: 
Structure of AP-2 bound to the dileucine motif and WxxPhi motif of CCDC32; consensus refinement
Method: single particle / : Baker RW, Kikkawa M, Sloan DE, Yanagisawa H

EMDB-72476: 
His-tagged Glutamine Synthetase on a Ni-NTA lipid monolayer grid
Method: single particle / : Baker RW, Strauss JD

EMDB-72471: 
His-tagged beta galactosidase (LacZ) on a Ni-NTA lipid monolayer grid
Method: single particle / : Baker RW, Strauss JD

EMDB-72472: 
Human nucleosome structure on Nickel-NTA lipid affinity grid (C2 refinement)
Method: single particle / : Baker RW, Strauss JD, McGinty RK, Skrajna A

EMDB-72473: 
Human nucleosome structure on Nickel-NTA lipid affinity grid (C1 refinement)
Method: single particle / : Baker RW, Strauss JD, McGinty RK, Skrajna A

EMDB-72474: 
Sro7 bound to His-Exo84 (1-326) on a Nickel-NTA lipid monolayer
Method: single particle / : Baker RW, Strauss JD, McGinty RK

EMDB-45207: 
AP-3 bound to myristoylated Arf1 (Q71L)
Method: single particle / : Begley MC, Baker RW

EMDB-45208: 
Human AP-3 dimer bound to myristoylated Arf1 (Q71L) and LAMP1 cargo on a lipid nanodisc
Method: single particle / : Begley MC, Baker RW

EMDB-45209: 
AP-3 Arf1 dimeric interface, focused refinement
Method: single particle / : Begley MC, Baker RW

EMDB-45210: 
AP-3 bound to myristoylated Arf1 and LAMPI on a lipid nanodisc; concensus refinement
Method: single particle / : Baker RW, Begley M

EMDB-45211: 
AP-3 bound to myristoylated Arf1 and LAMPI on a lipid nanodisc; focus refinement 1
Method: single particle / : Baker RW, Begley M

EMDB-45212: 
AP-3 bound to myristoylated Arf1 and LAMPI on a lipid nanodisc; focus refinement 2
Method: single particle / : Baker RW, Begley M

EMDB-45213: 
AP-3 bound to myristoylated Arf1 (Q71L) and LAMPI on a lipid nanodisc; combined map
Method: single particle / : Begley MC, Baker RW

EMDB-45214: 
Structure of Human Adaptor Protein Complex AP-3 in the Apo State
Method: single particle / : Begley MC, Baker RW

EMDB-16803: 
AP2 bound to MSP1 nanodisc with Tgn38 cargo peptide
Method: single particle / : Baker RW, Sarsam R, Cannon KS

EMDB-16804: 
AP2 bound to MSP1E3D1 nanodisc with Tgn38 cargo peptide
Method: single particle / : Baker RW, Sarsam R, Cannon KS

EMDB-29977: 
AP2 bound to an MSP1 nanodisc
Method: single particle / : Sarsam RD, Cannon KS, Baker RW

EMDB-40034: 
Structure of AP2 bound to MSP1E3D1 nanodisc
Method: single particle / : Sarsam RD, Cannon KS, Baker RW

EMDB-40035: 
Structure of AP2 bound to MSP2N2 nanodisc
Method: single particle / : Sarsam RD, Cannon KS, Baker RW

EMDB-40963: 
AP2 bound to MSP2N2 nanodisc with Tgn38 cargo peptide; focused refinement 1
Method: single particle / : Baker RW, Sarsam R, Cannon KS

EMDB-40964: 
AP2 bound to MSP2N2 nanodisc with Tgn38 cargo peptide; focused refinement 2
Method: single particle / : Baker RW, Sarsam R, Cannon KS

EMDB-40965: 
AP2 bound to MSP2N2 nanodisc with Tgn38 cargo peptide; focused refinement 3
Method: single particle / : Baker RW, Sarsam R, Cannon KS

EMDB-40966: 
AP2 bound to MSP2N2 nanodisc with Tgn38 cargo peptide; consensus refinement
Method: single particle / : Baker RW, Sarsam R, Cannon KS

EMDB-40973: 
AP2 bound to MSP2N2 nanodisc with Tgn38 cargo peptide; composite map
Method: single particle / : Baker RW, Cannon KS, Reta S

EMDB-24710: 
AP2 bound to heparin in the closed conformation
Method: single particle / : Baker RW, Hollopeter G

EMDB-24711: 
AP2 bound to heparin in the bowl conformation
Method: single particle / : Baker RW, Hollopeter G

EMDB-24712: 
AP2 bound to heparin and Tgn38 tyrosine cargo peptide
Method: single particle / : Baker RW, Hollopeter G

EMDB-24713: 
AP2 bound to the APA domain of SGIP in the presence of heparin
Method: single particle / : Baker RW, Hollopeter G

EMDB-24714: 
AP2 bound to the APA domain of SGIP and heparin; partial signal subtraction and symmetry expansion
Method: single particle / : Baker RW, Hollopeter G
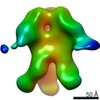
EMDB-23175: 
nsEM maps of BG505 SOSIP MD39 in complex with the polyclonal Fab samples from Group 1 of immunized rhesus macaques (Animal IDs: Rh.32034, Rh.32113, Rh.33104, Rh.33395, Rh.34909, Rh.34943)
Method: single particle / : Antanasijevic A, Sewall LM, Ward A
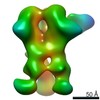
EMDB-23176: 
nsEM maps of BG505 SOSIP.v5.2 in complex with the polyclonal Fab samples from Group 2 of immunized rhesus macaques (Animal IDs: Rh.33182, Rh.33203, Rh.33311, Rh.34686, Rh.CD99, Rh.CG41)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB

EMDB-23177: 
nsEM maps of BG505 SOSIP.v5.2(7S) in complex with the polyclonal Fab samples from Group 3 of immunized rhesus macaques (Wk8 time point; Animal IDs: Rh.33172, Rh.34919, Rh.34167, Rh.33065, Rh.33176, Rh.34725)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB

EMDB-23178: 
nsEM maps of BG505 SOSIP.v5.2(7S) in complex with the polyclonal Fab samples from Group 3 of immunized rhesus macaques (Wk10 time point; Animal IDs: Rh.33172, Rh.34919, Rh.34167, Rh.33065, Rh.33176, Rh.34725)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB

EMDB-23179: 
nsEM maps of BG505 SOSIP.v5.2(7S) in complex with the polyclonal Fab samples from Group 3 of immunized rhesus macaques (Wk26 time point; Animal IDs: Rh.33172, Rh.34919, Rh.34167, Rh.33065, Rh.34725)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB

EMDB-23180: 
nsEM maps of BG505 SOSIP.v5.2(7S) in complex with the polyclonal Fab samples from Group 3 of immunized rhesus macaques (Wk38 time point; Animal IDs: Rh.33172, Rh.34919, Rh.34167, Rh.33065, Rh.34725)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB

EMDB-23181: 
nsEM map of BG505 SOSIP.v5.2(7S) in complex with the polyclonal Fab sample from animal Rh.33172 (Wk26 time point)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB

EMDB-23182: 
nsEM map of BG505 SOSIP.v3 in complex with the polyclonal Fab sample from animal Rh.33172 (Wk26 time point)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB
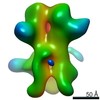
EMDB-23183: 
nsEM map of BG505 SOSIP.v3 in complex with the polyclonal Fab sample from animal Rh.33311 (Wk26 time point)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB
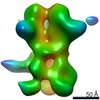
EMDB-23184: 
nsEM map of BG505 SOSIP.v5.2 in complex with the polyclonal Fab sample from animal Rh.33311 (Wk26 time point)
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB

EMDB-23185: 
BG505 SOSIP fused to a trimeric nanoparticle building block, T33-31A
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB
EMDB-23186: 
BG505 SOSIP fused to a trimeric nanoparticle building block, T33-31B
Method: single particle / : Antanasijevic A, Sewall LM, Ward AB

EMDB-23218: 
BG505 SOSIP reconstructed from a designed nanoparticle, BG505 SOSIP-T33-31 (Component A)
Method: single particle / : Antanasijevic A, Cannac F
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
