+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-8120 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of TRPV1 in complex with capsazepine, determined in lipid nanodisc | |||||||||||||||
 マップデータ マップデータ | None | |||||||||||||||
 試料 試料 |
| |||||||||||||||
| 生物種 |  | |||||||||||||||
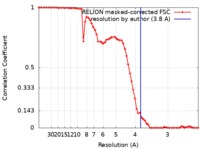
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.8 Å | |||||||||||||||
 データ登録者 データ登録者 | Gao Y / Cao E / Julius D / Cheng Y | |||||||||||||||
| 資金援助 |  米国, 4件 米国, 4件
| |||||||||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2016 ジャーナル: Nature / 年: 2016タイトル: TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. 著者: Yuan Gao / Erhu Cao / David Julius / Yifan Cheng /  要旨: When integral membrane proteins are visualized in detergents or other artificial systems, an important layer of information is lost regarding lipid interactions and their effects on protein structure. ...When integral membrane proteins are visualized in detergents or other artificial systems, an important layer of information is lost regarding lipid interactions and their effects on protein structure. This is especially relevant to proteins for which lipids have both structural and regulatory roles. Here we demonstrate the power of combining electron cryo-microscopy with lipid nanodisc technology to ascertain the structure of the rat TRPV1 ion channel in a native bilayer environment. Using this approach, we determined the locations of annular and regulatory lipids and showed that specific phospholipid interactions enhance binding of a spider toxin to TRPV1 through formation of a tripartite complex. Furthermore, phosphatidylinositol lipids occupy the binding site for capsaicin and other vanilloid ligands, suggesting a mechanism whereby chemical or thermal stimuli elicit channel activation by promoting the release of bioactive lipids from a critical allosteric regulatory site. | |||||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_8120.map.gz emd_8120.map.gz | 25 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-8120-v30.xml emd-8120-v30.xml emd-8120.xml emd-8120.xml | 21.5 KB 21.5 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_8120_fsc.xml emd_8120_fsc.xml | 6.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_8120.png emd_8120.png | 87.7 KB | ||
| その他 |  emd_8120_half_map_1.map.gz emd_8120_half_map_1.map.gz emd_8120_half_map_2.map.gz emd_8120_half_map_2.map.gz | 17.8 MB 17.8 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8120 http://ftp.pdbj.org/pub/emdb/structures/EMD-8120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8120 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8120 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_8120.map.gz / 形式: CCP4 / 大きさ: 27 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_8120.map.gz / 形式: CCP4 / 大きさ: 27 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.2156 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
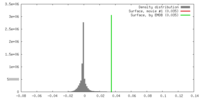
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-ハーフマップ: None
| ファイル | emd_8120_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | None | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
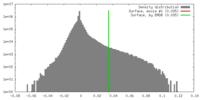
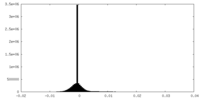
| 密度ヒストグラム |
-ハーフマップ: None
| ファイル | emd_8120_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | None | ||||||||||||
| 投影像・断面図 |
| ||||||||||||
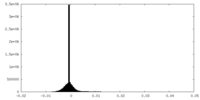
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
-全体 : TRPV1 ion channel in complex with capsazepine
| 全体 | 名称: TRPV1 ion channel in complex with capsazepine |
|---|---|
| 要素 |
|
-超分子 #1: TRPV1 ion channel in complex with capsazepine
| 超分子 | 名称: TRPV1 ion channel in complex with capsazepine / タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: all |
|---|---|
| 由来(天然) | 生物種:  |
| 組換発現 | 生物種:  Homo sapiens (ヒト) / 組換細胞: HEK 293S Homo sapiens (ヒト) / 組換細胞: HEK 293S |
-分子 #1: The capsaicin receptor, TRPV1
| 分子 | 名称: The capsaicin receptor, TRPV1 / タイプ: protein_or_peptide / ID: 1 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:  |
| 組換発現 | 生物種:  Homo sapiens (ヒト) Homo sapiens (ヒト) |
| 配列 | 文字列: AMGSRLYDRR SIFDAVAQSN CQELESLLPF LQRSKKRLTD SEFKDPETGK TCLLKAMLNL HNGQNDTIAL LLDVARKTDS LKQFVNASYT DSYYKGQTAL HIAIERRNMT LVTLLVENGA DVQAAANGDF FKKTKGRPGF YFGELPLSLA ACTNQLAIVK FLLQNSWQPA ...文字列: AMGSRLYDRR SIFDAVAQSN CQELESLLPF LQRSKKRLTD SEFKDPETGK TCLLKAMLNL HNGQNDTIAL LLDVARKTDS LKQFVNASYT DSYYKGQTAL HIAIERRNMT LVTLLVENGA DVQAAANGDF FKKTKGRPGF YFGELPLSLA ACTNQLAIVK FLLQNSWQPA DISARDSVGN TVLHALVEVA DNTVDNTKFV TSMYNEILIL GAKLHPTLKL EEITNRKGLT PLALAASSGK IGVLAYILQR EIHEPECRHL SRKFTEWAYG PVHSSLYDLS CIDTCEKNSV LEVIAYSSSE TPNRHDMLLV EPLNRLLQDK WDRFVKRIFY FNFFVYCLYM IIFTAAAYYR PVEGLPPYKL KNTVGDYFRV TGEILSVSGG VYFFFRGIQY FLQRRPSLKS LFVDSYSEIL FFVQSLFMLV SVVLYFSQRK EYVASMVFSL AMGWTNMLYY TRGFQQMGIY AVMIEKMILR DLCRFMFVYL VFLFGFSTAV VTLIEDGKYN SLYSTCLELF KFTIGMGDLE FTENYDFKAV FIILLLAYVI LTYILLLNML IALMGETVNK IAQESKNIWK LQRAITILDT EKSFLKCMRK AFRSGKLLQV GFTPDGKDDY RWCFRVDEVN WTTWNTNVGI INEDPG |
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 7.4 構成要素:
| ||||||||||||
| グリッド | モデル: Quantifoil R1.2/1.3 / 材質: COPPER / メッシュ: 400 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 支持フィルム - Film thickness: 12.0 nm / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 雰囲気: AIR / 前処理 - 気圧: 290.0 kPa | ||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 298 K / 装置: FEI VITROBOT MARK III 詳細: Blot for 6 seconds before plunging into liquid ethane (FEI VITROBOT MARK III).. | ||||||||||||
| 詳細 | This sample was monodisperse. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI POLARA 300 |
|---|---|
| 温度 | 最低: 70.0 K / 最高: 70.0 K |
| 詳細 | Grid screening was performed manually. |
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: SUPER-RESOLUTION / デジタル化 - サイズ - 横: 3838 pixel / デジタル化 - サイズ - 縦: 3710 pixel / デジタル化 - 画像ごとのフレーム数: 1-30 / 撮影したグリッド数: 1 / 実像数: 1002 / 平均露光時間: 6.0 sec. / 平均電子線量: 41.0 e/Å2 詳細: Images were collected in movie mode at 5 frames per second for 6 seconds. |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 30.0 µm / 最大 デフォーカス(補正後): 2.2 µm / 最小 デフォーカス(補正後): 0.7 µm / 倍率(補正後): 41132 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 2.0 mm / 最大 デフォーカス(公称値): 2.2 µm / 最小 デフォーカス(公称値): 0.7 µm / 倍率(公称値): 31000 |
| 試料ステージ | 試料ホルダーモデル: OTHER / ホルダー冷却材: NITROGEN |
| 実験機器 |  モデル: Tecnai Polara / 画像提供: FEI Company |
+ 画像解析
画像解析
-原子モデル構築 1
| 初期モデル | PDB ID: Chain - Chain ID: A |
|---|---|
| 精密化 | 空間: REAL / プロトコル: RIGID BODY FIT / 当てはまり具合の基準: correlation coefficient |
 ムービー
ムービー コントローラー
コントローラー

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)