+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8118 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of TRPV1 determined in lipid nanodisc | |||||||||||||||
 Map data Map data | TRPV1 | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | TRP / ion channel / nanodisc / vanilloid / lipid / TRANSPORT PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of iodide transmembrane transport / positive regulation of membrane depolarization / negative regulation of establishment of blood-brain barrier / response to capsazepine / sensory perception of mechanical stimulus / peptide secretion / cellular response to temperature stimulus / positive regulation of sensory perception of pain / temperature-gated ion channel activity / detection of chemical stimulus involved in sensory perception of pain ...negative regulation of iodide transmembrane transport / positive regulation of membrane depolarization / negative regulation of establishment of blood-brain barrier / response to capsazepine / sensory perception of mechanical stimulus / peptide secretion / cellular response to temperature stimulus / positive regulation of sensory perception of pain / temperature-gated ion channel activity / detection of chemical stimulus involved in sensory perception of pain / positive regulation of renal sodium excretion / TRP channels / negative regulation of axon regeneration / positive regulation of cardiac muscle cell differentiation / smooth muscle contraction involved in micturition / fever generation / detection of temperature stimulus involved in thermoception / thermoception / urinary bladder smooth muscle contraction / response to pH / glutamate secretion / monoatomic cation transmembrane transporter activity / negative regulation of systemic arterial blood pressure / excitatory extracellular ligand-gated monoatomic ion channel activity / dendritic spine membrane / negative regulation of heart rate / response to acidic pH / positive regulation of urine volume / response to pain / diet induced thermogenesis / cellular response to alkaloid / temperature homeostasis / cellular response to cytokine stimulus / cellular response to ATP / intracellularly gated calcium channel activity / detection of temperature stimulus involved in sensory perception of pain / negative regulation of mitochondrial membrane potential / calcium ion import across plasma membrane / behavioral response to pain / positive regulation of vasoconstriction / monoatomic ion channel activity / ligand-gated monoatomic ion channel activity / monoatomic cation channel activity / cellular response to acidic pH / extracellular ligand-gated monoatomic ion channel activity / phosphatidylinositol binding / sensory perception of pain / axon terminus / positive regulation of excitatory postsynaptic potential / sarcoplasmic reticulum / lipid metabolic process / phosphoprotein binding / microglial cell activation / cellular response to nerve growth factor stimulus / response to peptide hormone / cellular response to growth factor stimulus / GABA-ergic synapse / cellular response to tumor necrosis factor / calcium ion transmembrane transport / calcium channel activity / positive regulation of nitric oxide biosynthetic process / calcium ion transport / transmembrane signaling receptor activity / cellular response to heat / sensory perception of taste / response to heat / positive regulation of cytosolic calcium ion concentration / monoatomic ion transmembrane transport / protein homotetramerization / calmodulin binding / postsynaptic membrane / neuron projection / positive regulation of apoptotic process / external side of plasma membrane / neuronal cell body / dendrite / negative regulation of transcription by RNA polymerase II / ATP binding / membrane / metal ion binding / identical protein binding / nucleus / plasma membrane Similarity search - Function | |||||||||||||||
| Biological species |  | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.28 Å | |||||||||||||||
 Authors Authors | Gao Y / Cao E | |||||||||||||||
| Funding support |  United States, 4 items United States, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nature / Year: 2016 Journal: Nature / Year: 2016Title: TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Authors: Yuan Gao / Erhu Cao / David Julius / Yifan Cheng /  Abstract: When integral membrane proteins are visualized in detergents or other artificial systems, an important layer of information is lost regarding lipid interactions and their effects on protein structure. ...When integral membrane proteins are visualized in detergents or other artificial systems, an important layer of information is lost regarding lipid interactions and their effects on protein structure. This is especially relevant to proteins for which lipids have both structural and regulatory roles. Here we demonstrate the power of combining electron cryo-microscopy with lipid nanodisc technology to ascertain the structure of the rat TRPV1 ion channel in a native bilayer environment. Using this approach, we determined the locations of annular and regulatory lipids and showed that specific phospholipid interactions enhance binding of a spider toxin to TRPV1 through formation of a tripartite complex. Furthermore, phosphatidylinositol lipids occupy the binding site for capsaicin and other vanilloid ligands, suggesting a mechanism whereby chemical or thermal stimuli elicit channel activation by promoting the release of bioactive lipids from a critical allosteric regulatory site. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8118.map.gz emd_8118.map.gz | 24.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8118-v30.xml emd-8118-v30.xml emd-8118.xml emd-8118.xml | 23.2 KB 23.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8118_fsc.xml emd_8118_fsc.xml | 6.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_8118.png emd_8118.png | 210.3 KB | ||
| Filedesc metadata |  emd-8118.cif.gz emd-8118.cif.gz | 7.2 KB | ||
| Others |  emd_8118_half_map_1.map.gz emd_8118_half_map_1.map.gz emd_8118_half_map_2.map.gz emd_8118_half_map_2.map.gz | 17.9 MB 17.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8118 http://ftp.pdbj.org/pub/emdb/structures/EMD-8118 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8118 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8118 | HTTPS FTP |
-Related structure data
| Related structure data |  5irzMC  8117C  8119C  8120C  5irxC  5is0C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8118.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8118.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TRPV1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.2156 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: TRPV1, half map 1
| File | emd_8118_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TRPV1, half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: TRPV1, half map 2
| File | emd_8118_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | TRPV1, half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : TRPV1 ion channel in unliganded state
| Entire | Name: TRPV1 ion channel in unliganded state |
|---|---|
| Components |
|
-Supramolecule #1: TRPV1 ion channel in unliganded state
| Supramolecule | Name: TRPV1 ion channel in unliganded state / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Transient receptor potential cation channel subfamily V member 1
| Macromolecule | Name: Transient receptor potential cation channel subfamily V member 1 type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 72.959297 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AMGSRLYDRR SIFDAVAQSN CQELESLLPF LQRSKKRLTD SEFKDPETGK TCLLKAMLNL HNGQNDTIAL LLDVARKTDS LKQFVNASY TDSYYKGQTA LHIAIERRNM TLVTLLVENG ADVQAAANGD FFKKTKGRPG FYFGELPLSL AACTNQLAIV K FLLQNSWQ ...String: AMGSRLYDRR SIFDAVAQSN CQELESLLPF LQRSKKRLTD SEFKDPETGK TCLLKAMLNL HNGQNDTIAL LLDVARKTDS LKQFVNASY TDSYYKGQTA LHIAIERRNM TLVTLLVENG ADVQAAANGD FFKKTKGRPG FYFGELPLSL AACTNQLAIV K FLLQNSWQ PADISARDSV GNTVLHALVE VADNTVDNTK FVTSMYNEIL ILGAKLHPTL KLEEITNRKG LTPLALAASS GK IGVLAYI LQREIHEPEC RHLSRKFTEW AYGPVHSSLY DLSCIDTCEK NSVLEVIAYS SSETPNRHDM LLVEPLNRLL QDK WDRFVK RIFYFNFFVY CLYMIIFTAA AYYRPVEGLP PYKLKNTVGD YFRVTGEILS VSGGVYFFFR GIQYFLQRRP SLKS LFVDS YSEILFFVQS LFMLVSVVLY FSQRKEYVAS MVFSLAMGWT NMLYYTRGFQ QMGIYAVMIE KMILRDLCRF MFVYL VFLF GFSTAVVTLI EDGKYNSLYS TCLELFKFTI GMGDLEFTEN YDFKAVFIIL LLAYVILTYI LLLNMLIALM GETVNK IAQ ESKNIWKLQR AITILDTEKS FLKCMRKAFR SGKLLQVGFT PDGKDDYRWC FRVDEVNWTT WNTNVGIINE DPG UniProtKB: Transient receptor potential cation channel subfamily V member 1, Transient receptor potential cation channel subfamily V member 1 |
-Macromolecule #2: (4R,7S)-4-hydroxy-N,N,N-trimethyl-4,9-dioxo-7-[(pentanoyloxy)meth...
| Macromolecule | Name: (4R,7S)-4-hydroxy-N,N,N-trimethyl-4,9-dioxo-7-[(pentanoyloxy)methyl]-3,5,8-trioxa-4lambda~5~-phosphatetradecan-1-aminium type: ligand / ID: 2 / Number of copies: 4 / Formula: 6O8 |
|---|---|
| Molecular weight | Theoretical: 440.489 Da |
| Chemical component information | 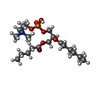 ChemComp-6O8: |
-Macromolecule #3: (2S)-1-{[(R)-hydroxy{[(1R,2R,3S,4S,5S,6S)-2,3,4,5,6-pentahydroxyc...
| Macromolecule | Name: (2S)-1-{[(R)-hydroxy{[(1R,2R,3S,4S,5S,6S)-2,3,4,5,6-pentahydroxycyclohexyl]oxy}phosphoryl]oxy}-3-(pentanoyloxy)propan-2-yl decanoate type: ligand / ID: 3 / Number of copies: 4 / Formula: 6ES |
|---|---|
| Molecular weight | Theoretical: 572.58 Da |
| Chemical component information | 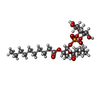 ChemComp-6ES: |
-Macromolecule #4: (2S)-3-{[(S)-(2-aminoethoxy)(hydroxy)phosphoryl]oxy}-2-(hexanoylo...
| Macromolecule | Name: (2S)-3-{[(S)-(2-aminoethoxy)(hydroxy)phosphoryl]oxy}-2-(hexanoyloxy)propyl hexanoate type: ligand / ID: 4 / Number of copies: 16 / Formula: 6OE |
|---|---|
| Molecular weight | Theoretical: 411.428 Da |
| Chemical component information | 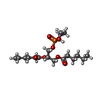 ChemComp-6OE: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 12 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 290.0 kPa | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 278 K / Instrument: FEI VITROBOT MARK III Details: Blot for 6 seconds before plunging into liquid ethane (FEI VITROBOT MARK III).. | ||||||||||||
| Details | This sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Temperature | Min: 70.0 K / Max: 70.0 K |
| Details | Grid screening was performed manually. |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Frames/image: 1-30 / Number grids imaged: 1 / Number real images: 1000 / Average exposure time: 6.0 sec. / Average electron dose: 41.0 e/Å2 Details: Images were collected in movie mode at 5 frames per second for 6 seconds. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 30.0 µm / Calibrated defocus max: 2.2 µm / Calibrated defocus min: 0.7000000000000001 µm / Calibrated magnification: 41132 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 31000 |
| Sample stage | Specimen holder model: OTHER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)







































