[English] 日本語
 Yorodumi
Yorodumi- EMDB-12804: Resting state GluA1/A2 heterotetramer in complex with auxiliary s... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12804 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Resting state GluA1/A2 heterotetramer in complex with auxiliary subunit TARP gamma 8 (LBD-TMD) | |||||||||
 Map data Map data | Composite map generated based on masked refinement maps of LBD and TMD | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AMPAR / ion channels / neurotransmission / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationPhase 0 - rapid depolarisation / Phase 2 - plateau phase / Cargo concentration in the ER / axonal spine / positive regulation of locomotion involved in locomotory behavior / COPII-mediated vesicle transport / positive regulation of membrane potential / cellular response to ammonium ion / response to sucrose / L-type voltage-gated calcium channel complex ...Phase 0 - rapid depolarisation / Phase 2 - plateau phase / Cargo concentration in the ER / axonal spine / positive regulation of locomotion involved in locomotory behavior / COPII-mediated vesicle transport / positive regulation of membrane potential / cellular response to ammonium ion / response to sucrose / L-type voltage-gated calcium channel complex / myosin V binding / LGI-ADAM interactions / neuron spine / Trafficking of AMPA receptors / postsynaptic neurotransmitter receptor diffusion trapping / proximal dendrite / regulation of AMPA receptor activity / protein phosphatase 2B binding / response to arsenic-containing substance / regulation of monoatomic ion transmembrane transport / channel regulator activity / cellular response to L-glutamate / cellular response to dsRNA / ligand-gated calcium channel activity / dendritic spine membrane / Synaptic adhesion-like molecules / beta-2 adrenergic receptor binding / long-term synaptic depression / cellular response to peptide hormone stimulus / spine synapse / dendritic spine neck / dendritic spine cytoplasm / dendritic spine head / cellular response to amine stimulus / peptide hormone receptor binding / response to psychosocial stress / response to morphine / Activation of AMPA receptors / ligand-gated monoatomic cation channel activity / spinal cord development / perisynaptic space / neuronal cell body membrane / protein kinase A binding / Trafficking of GluR2-containing AMPA receptors / response to lithium ion / AMPA glutamate receptor activity / AMPA glutamate receptor clustering / transmission of nerve impulse / kainate selective glutamate receptor activity / behavioral response to pain / immunoglobulin binding / adenylate cyclase binding / asymmetric synapse / response to electrical stimulus / extracellularly glutamate-gated ion channel activity / AMPA glutamate receptor complex / cellular response to glycine / regulation of receptor recycling / ionotropic glutamate receptor complex / Unblocking of NMDA receptors, glutamate binding and activation / G-protein alpha-subunit binding / glutamate receptor binding / conditioned place preference / positive regulation of synaptic transmission / long-term memory / regulation of postsynaptic membrane neurotransmitter receptor levels / voltage-gated calcium channel activity / postsynaptic density, intracellular component / response to fungicide / regulation of synaptic transmission, glutamatergic / neuronal action potential / positive regulation of synaptic transmission, glutamatergic / extracellular ligand-gated monoatomic ion channel activity / cytoskeletal protein binding / glutamate-gated receptor activity / regulation of long-term synaptic depression / cellular response to brain-derived neurotrophic factor stimulus / somatodendritic compartment / glutamate-gated calcium ion channel activity / presynaptic active zone membrane / synapse assembly / excitatory synapse / ionotropic glutamate receptor signaling pathway / ionotropic glutamate receptor binding / dendrite membrane / dendrite cytoplasm / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / positive regulation of excitatory postsynaptic potential / dendritic shaft / SNARE binding / synaptic membrane / response to cocaine / PDZ domain binding / calcium channel regulator activity / neuromuscular junction / cellular response to amino acid stimulus / synaptic transmission, glutamatergic / protein tetramerization / establishment of protein localization / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential Similarity search - Function | |||||||||
| Biological species |  | |||||||||
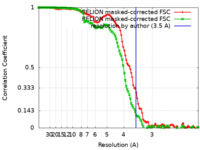
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Zhang D / Watson JF / Matthews PM / Cais O / Greger IH | |||||||||
| Funding support | 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2021 Journal: Nature / Year: 2021Title: Gating and modulation of a hetero-octameric AMPA glutamate receptor. Authors: Danyang Zhang / Jake F Watson / Peter M Matthews / Ondrej Cais / Ingo H Greger /   Abstract: AMPA receptors (AMPARs) mediate the majority of excitatory transmission in the brain and enable the synaptic plasticity that underlies learning. A diverse array of AMPAR signalling complexes are ...AMPA receptors (AMPARs) mediate the majority of excitatory transmission in the brain and enable the synaptic plasticity that underlies learning. A diverse array of AMPAR signalling complexes are established by receptor auxiliary subunits, which associate with the AMPAR in various combinations to modulate trafficking, gating and synaptic strength. However, their mechanisms of action are poorly understood. Here we determine cryo-electron microscopy structures of the heteromeric GluA1-GluA2 receptor assembled with both TARP-γ8 and CNIH2, the predominant AMPAR complex in the forebrain, in both resting and active states. Two TARP-γ8 and two CNIH2 subunits insert at distinct sites beneath the ligand-binding domains of the receptor, with site-specific lipids shaping each interaction and affecting the gating regulation of the AMPARs. Activation of the receptor leads to asymmetry between GluA1 and GluA2 along the ion conduction path and an outward expansion of the channel triggers counter-rotations of both auxiliary subunit pairs, promoting the active-state conformation. In addition, both TARP-γ8 and CNIH2 pivot towards the pore exit upon activation, extending their reach for cytoplasmic receptor elements. CNIH2 achieves this through its uniquely extended M2 helix, which has transformed this endoplasmic reticulum-export factor into a powerful AMPAR modulator that is capable of providing hippocampal pyramidal neurons with their integrative synaptic properties. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12804.map.gz emd_12804.map.gz | 6.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12804-v30.xml emd-12804-v30.xml emd-12804.xml emd-12804.xml | 20.7 KB 20.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12804_fsc.xml emd_12804_fsc.xml emd_12804_fsc_2.xml emd_12804_fsc_2.xml | 11.4 KB 11.4 KB | Display Display |  FSC data file FSC data file |
| Images |  emd_12804.png emd_12804.png | 171.8 KB | ||
| Filedesc metadata |  emd-12804.cif.gz emd-12804.cif.gz | 7.1 KB | ||
| Others |  emd_12804_additional_1.map.gz emd_12804_additional_1.map.gz emd_12804_additional_2.map.gz emd_12804_additional_2.map.gz | 3.6 MB 5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12804 http://ftp.pdbj.org/pub/emdb/structures/EMD-12804 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12804 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12804 | HTTPS FTP |
-Related structure data
| Related structure data |  7ocdMC  7ocaC  7occC  7oceC  7ocfC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12804.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12804.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Composite map generated based on masked refinement maps of LBD and TMD | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: Masked refinement map focusing on LBD
| File | emd_12804_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Masked refinement map focusing on LBD | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Masked refinement map focusing on TMD
| File | emd_12804_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Masked refinement map focusing on TMD | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : GluA1/A2 heterotetramer in complex with auxiliary subunit TARP gamma 8
| Entire | Name: GluA1/A2 heterotetramer in complex with auxiliary subunit TARP gamma 8 |
|---|---|
| Components |
|
-Supramolecule #1: GluA1/A2 heterotetramer in complex with auxiliary subunit TARP gamma 8
| Supramolecule | Name: GluA1/A2 heterotetramer in complex with auxiliary subunit TARP gamma 8 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Isoform Flip of Glutamate receptor 1
| Macromolecule | Name: Isoform Flip of Glutamate receptor 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 102.66193 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPYIFAFFCT GFLGAVVGAD YKDDDDKNFP NNIQIGGLFP NQQSQEHAAF RFALSQLTEP PKLLPQIDIV NISDSFEMTY RFCSQFSKG VYAIFGFYER RTVNMLTSFC GALHVCFITP SFPVDTSNQF VLQLRPELQE ALISIIDHYK WQTFVYIYDA D RGLSVLQR ...String: MPYIFAFFCT GFLGAVVGAD YKDDDDKNFP NNIQIGGLFP NQQSQEHAAF RFALSQLTEP PKLLPQIDIV NISDSFEMTY RFCSQFSKG VYAIFGFYER RTVNMLTSFC GALHVCFITP SFPVDTSNQF VLQLRPELQE ALISIIDHYK WQTFVYIYDA D RGLSVLQR VLDTAAEKNW QVTAVNILTT TEEGYRMLFQ DLEKKKERLV VVDCESERLN AILGQIVKLE KNGIGYHYIL AN LGFMDID LNKFKESGAN VTGFQLVNYT DTIPARIMQQ WRTSDSRDHT RVDWKRPKYT SALTYDGVKV MAEAFQSLRR QRI DISRRG NAGDCLANPA VPWGQGIDIQ RALQQVRFEG LTGNVQFNEK GRRTNYTLHV IEMKHDGIRK IGYWNEDDKF VPAA TDAQA GGDNSSVQNR TYIVTTILED PYVMLKKNAN QFEGNDRYEG YCVELAAEIA KHVGYSYRLE IVSDGKYGAR DPDTK AWNG MVGELVYGRA DVAVAPLTIT LVREEVIDFS KPFMSLGISI MIKKPQKSKP GVFSFLDPLA YEIWMCIVFA YIGVSV VLF LVSRFSPYEW HSEEFEEGRD QTTSDQSNEF GIFNSLWFSL GAFMQQGCDI SPRSLSGRIV GGVWWFFTLI IISSYTA NL AAFLTVERMV SPIESAEDLA KQTEIAYGTL EAGSTKEFFR RSKIAVFEKM WTYMKSAEPS VFVRTTEEGM IRVRKSKG K YAYLLESTMN EYIEQRKPCD TMKVGGNLDS KGYGIATPKG SALRGPVNLA VLKLSEQGVL DKLKSKWWYD KGECGSKDS GSKDKTSALS LSNVAGVFYI LIGGLGLAML VALIEFCYKS RSESKRMKGF CLIPQQSINE AIRTSTLPRN SGAGASGGGG SGENGRVVS QDFPKSMQSI PCMSHSSGMP LGATGL UniProtKB: Glutamate receptor 1 |
-Macromolecule #2: Glutamate receptor 2
| Macromolecule | Name: Glutamate receptor 2 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 96.247055 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MQKIMHISVL LSPVLWGLIF GVSSNSIQIG GLFPRGADQE YSAFRVGMVQ FSTSEFRLTP HIDNLEVANS FAVTNAFCSQ FSRGVYAIF GFYDKKSVNT ITSFCGTLHV SFITPSFPTD GTHPFVIQMR PDLKGALLSL IEYYQWDKFA YLYDSDRGLS T LQAVLDSA ...String: MQKIMHISVL LSPVLWGLIF GVSSNSIQIG GLFPRGADQE YSAFRVGMVQ FSTSEFRLTP HIDNLEVANS FAVTNAFCSQ FSRGVYAIF GFYDKKSVNT ITSFCGTLHV SFITPSFPTD GTHPFVIQMR PDLKGALLSL IEYYQWDKFA YLYDSDRGLS T LQAVLDSA AEKKWQVTAI NVGNINNDKK DETYRSLFQD LELKKERRVI LDCERDKVND IVDQVITIGK HVKGYHYIIA NL GFTDGDL LKIQFGGANV SGFQIVDYDD SLVSKFIERW STLEEKEYPG AHTATIKYTS ALTYDAVQVM TEAFRNLRKQ RIE ISRRGN AGDCLANPAV PWGQGVEIER ALKQVQVEGL SGNIKFDQNG KRINYTINIM ELKTNGPRKI GYWSEVDKMV VTLT ELPSG NDTSGLENKT VVVTTILESP YVMMKKNHEM LEGNERYEGY CVDLAAEIAK HCGFKYKLTI VGDGKYGARD ADTKI WNGM VGELVYGKAD IAIAPLTITL VREEVIDFSK PFMSLGISIM IKKPQKSKPG VFSFLDPLAY EIWMCIVFAY IGVSVV LFL VSRFSPYEWH TEEFEDGRET QSSESTNEFG IFNSLWFSLG AFMRQGCDIS PRSLSGRIVG GVWWFFTLII ISSYTAN LA AFLTVERMVS PIESAEDLSK QTEIAYGTLD SGSTKEFFRR SKIAVFDKMW TYMRSAEPSV FVRTTAEGVA RVRKSKGK Y AYLLESTMNE YIEQRKPCDT MKVGGNLDSK GYGIATPKGS SLGTPVNLAV LKLSEQGVLD KLKNKWWYDK GECGAKDSG SKEKTSALSL SNVAGVFYIL VGGLGLAMLV ALIEFCYKSR AEAKRMKVAK NPQNINPSSS UniProtKB: Glutamate receptor 2 |
-Macromolecule #3: Voltage-dependent calcium channel gamma-8 subunit
| Macromolecule | Name: Voltage-dependent calcium channel gamma-8 subunit / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 43.576004 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GESLKRWNEE RGLWCEKGVQ VLLTTIGAFA AFGLMTIAIS TDYWLYTRAL ICNTTNLTAG DDGPPHRGGS GSSEKKDPGG LTHSGLWRI CCLEGLKRGV CVKINHFPED TDYDHDSAEY LLRVVRASSI FPILSAILLL LGGVCVAASR VYKSKRNIIL G AGILFVAA ...String: GESLKRWNEE RGLWCEKGVQ VLLTTIGAFA AFGLMTIAIS TDYWLYTRAL ICNTTNLTAG DDGPPHRGGS GSSEKKDPGG LTHSGLWRI CCLEGLKRGV CVKINHFPED TDYDHDSAEY LLRVVRASSI FPILSAILLL LGGVCVAASR VYKSKRNIIL G AGILFVAA GLSNIIGVIV YISANAGEPG PKRDEEKKNH YSYGWSFYFG GLSFILAEVI GVLAVNIYIE RSREAHCQSR SD LLKAGGG AGGSGGSGPS AILRLPSYRF RYRRRSRSSS RGSSEASPSR DASPGGPGGP GFASTDISMY TLSRDPSKGS VAA GLASAG GGGGGAGVGA YGGAAGAAGG GGTGSERDRG SSAGFLTLHN AFPKEAASGV TVTVTGPPAA PAPAPPAPAA PAPG TLSKE AAASNTNTLN RKLEVLFQ UniProtKB: Voltage-dependent calcium channel gamma-8 subunit |
-Macromolecule #4: 6-nitro-2,3-bis(oxidanylidene)-1,4-dihydrobenzo[f]quinoxaline-7-s...
| Macromolecule | Name: 6-nitro-2,3-bis(oxidanylidene)-1,4-dihydrobenzo[f]quinoxaline-7-sulfonamide type: ligand / ID: 4 / Number of copies: 4 / Formula: E2Q |
|---|---|
| Molecular weight | Theoretical: 336.28 Da |
| Chemical component information | 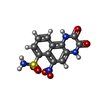 ChemComp-E2Q: |
-Macromolecule #5: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 5 / Number of copies: 6 / Formula: PC1 |
|---|---|
| Molecular weight | Theoretical: 790.145 Da |
| Chemical component information |  ChemComp-PC1: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
| Details | Purified protein was incubated with 100 uM NBQX for at least 30 min on ice before freezing. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)