+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10702 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Post-fusion X-31 Influenza Haemagglutinin at pH 5 (State V) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |   unidentified influenza virus unidentified influenza virus | |||||||||
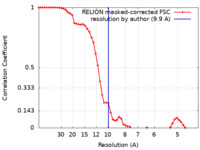
| Method | single particle reconstruction / cryo EM / Resolution: 9.9 Å | |||||||||
 Authors Authors | Benton DJ / Rosenthal PB | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2020 Journal: Nature / Year: 2020Title: Structural transitions in influenza haemagglutinin at membrane fusion pH. Authors: Donald J Benton / Steven J Gamblin / Peter B Rosenthal / John J Skehel /  Abstract: Infection by enveloped viruses involves fusion of their lipid envelopes with cellular membranes to release the viral genome into cells. For HIV, Ebola, influenza and numerous other viruses, envelope ...Infection by enveloped viruses involves fusion of their lipid envelopes with cellular membranes to release the viral genome into cells. For HIV, Ebola, influenza and numerous other viruses, envelope glycoproteins bind the infecting virion to cell-surface receptors and mediate membrane fusion. In the case of influenza, the receptor-binding glycoprotein is the haemagglutinin (HA), and following receptor-mediated uptake of the bound virus by endocytosis, it is the HA that mediates fusion of the virus envelope with the membrane of the endosome. Each subunit of the trimeric HA consists of two disulfide-linked polypeptides, HA1 and HA2. The larger, virus-membrane-distal, HA1 mediates receptor binding; the smaller, membrane-proximal, HA2 anchors HA in the envelope and contains the fusion peptide, a region that is directly involved in membrane interaction. The low pH of endosomes activates fusion by facilitating irreversible conformational changes in the glycoprotein. The structures of the initial HA at neutral pH and the final HA at fusion pH have been investigated by electron microscopy and X-ray crystallography. Here, to further study the process of fusion, we incubate HA for different times at pH 5.0 and directly image structural changes using single-particle cryo-electron microscopy. We describe three distinct, previously undescribed forms of HA, most notably a 150 Å-long triple-helical coil of HA2, which may bridge between the viral and endosomal membranes. Comparison of these structures reveals concerted conformational rearrangements through which the HA mediates membrane fusion. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10702.map.gz emd_10702.map.gz | 11.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10702-v30.xml emd-10702-v30.xml emd-10702.xml emd-10702.xml | 14.9 KB 14.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10702_fsc.xml emd_10702_fsc.xml | 5.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_10702.png emd_10702.png | 41.6 KB | ||
| Masks |  emd_10702_msk_1.map emd_10702_msk_1.map | 12.9 MB |  Mask map Mask map | |
| Others |  emd_10702_half_map_1.map.gz emd_10702_half_map_1.map.gz emd_10702_half_map_2.map.gz emd_10702_half_map_2.map.gz | 9.8 MB 9.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10702 http://ftp.pdbj.org/pub/emdb/structures/EMD-10702 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10702 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10702 | HTTPS FTP |
-Validation report
| Summary document |  emd_10702_validation.pdf.gz emd_10702_validation.pdf.gz | 288.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10702_full_validation.pdf.gz emd_10702_full_validation.pdf.gz | 287.5 KB | Display | |
| Data in XML |  emd_10702_validation.xml.gz emd_10702_validation.xml.gz | 10.8 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10702 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10702 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10702 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10702 | HTTPS FTP |
-Related structure data
| Related structure data |  6y5gC  6y5hC  6y5iC  6y5jC  6y5kC  6y5lC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10702.map.gz / Format: CCP4 / Size: 12.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10702.map.gz / Format: CCP4 / Size: 12.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.18 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10702_msk_1.map emd_10702_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
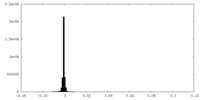
| Density Histograms |
-Half map: #1
| File | emd_10702_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_10702_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : X-31 Influenza Haemagglutinin
| Entire | Name: X-31 Influenza Haemagglutinin |
|---|---|
| Components |
|
-Supramolecule #1: X-31 Influenza Haemagglutinin
| Supramolecule | Name: X-31 Influenza Haemagglutinin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   unidentified influenza virus unidentified influenza virus |
| Recombinant expression | Organism:  |
| Molecular weight | Theoretical: 60 KDa |
-Macromolecule #1: X-31 Influenza Haemagglutinin HA2
| Macromolecule | Name: X-31 Influenza Haemagglutinin HA2 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   unidentified influenza virus unidentified influenza virus |
| Sequence | String: GLFGAIAGFI ENGWEGMIDG WYGFRHQNSE GTGQAADLKS TQAAIDQING KLNRVIEKTN EKFHQIEKEF SEVEGRIQDL EKYVEDTKID LWSYNAELLV ALENQHTIDL TDSEMNKLFE KTRRQLRENA EEMGNGCFKI YHKCDNACIE SIRNGTYDHD VYRDEALNNR FQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.5 mg/mL |
|---|---|
| Buffer | pH: 5 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Details: 25mA |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Average exposure time: 60.0 sec. / Average electron dose: 33.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)