1C6V
 
 | | SIV INTEGRASE (CATALYTIC DOMAIN + DNA BIDING DOMAIN COMPRISING RESIDUES 50-293) MUTANT WITH PHE 185 REPLACED BY HIS (F185H) | | Descriptor: | PROTEIN (SIU89134), PROTEIN (SIV INTEGRASE) | | Authors: | Chen, Z, Yan, Y, Munshi, S, Li, Y, Zruygay-Murphy, J, Xu, B, Witmer, M, Felock, P, Wolfe, A, Sardana, V, Emini, E.A, Hazuda, D, Kuo, L.C. | | Deposit date: | 1999-12-21 | | Release date: | 2000-12-27 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | X-ray structure of simian immunodeficiency virus integrase containing the core and C-terminal domain (residues 50-293)--an initial glance of the viral DNA binding platform.
J.Mol.Biol., 296, 2000
|
|
5W45
 
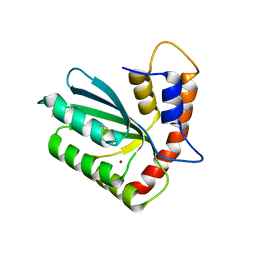 | | Crystal structure of APOBEC3H | | Descriptor: | APOBEC3H, ZINC ION | | Authors: | Ito, F, Yang, H.J, Xiao, X, Li, S.X, Wolfe, A, Zirkle, B, Arutiunian, V, Chen, X.S. | | Deposit date: | 2017-06-09 | | Release date: | 2018-03-14 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.486 Å) | | Cite: | Understanding the Structure, Multimerization, Subcellular Localization and mC Selectivity of a Genomic Mutator and Anti-HIV Factor APOBEC3H.
Sci Rep, 8, 2018
|
|
5W2M
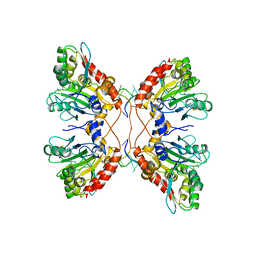 
 | | APOBEC3F Catalytic Domain Complex with a Single-Stranded DNA | | Descriptor: | DNA (5'-D(P*TP*TP*TP*TP*TP*TP*TP*TP*TP*T)-3'), DNA dC->dU-editing enzyme APOBEC-3F, ZINC ION | | Authors: | Fang, Y, Xiao, X, Li, S.-X, Wolfe, A, Chen, X.S. | | Deposit date: | 2017-06-06 | | Release date: | 2017-12-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Molecular Interactions of a DNA Modifying Enzyme APOBEC3F Catalytic Domain with a Single-Stranded DNA.
J. Mol. Biol., 430, 2018
|
|
2L2Z
 
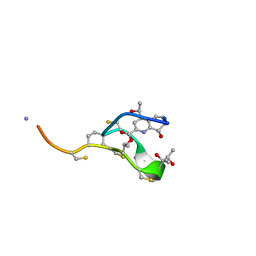 | | Thiostrepton, reduced at N-CA bond of residue 14 | | Descriptor: | Thiostrepton | | Authors: | Jonker, H.R.A, Baumann, S, Wolf, A, Schoof, S, Hiller, F, Schulte, K.W, Kirschner, K.N, Schwalbe, H, Arndt, H.-D. | | Deposit date: | 2010-08-27 | | Release date: | 2011-02-02 | | Last modified: | 2013-06-26 | | Method: | SOLUTION NMR | | Cite: | NMR structures of thiostrepton derivatives for characterization of the ribosomal binding site.
Angew.Chem.Int.Ed.Engl., 50, 2011
|
|
2L2W
 
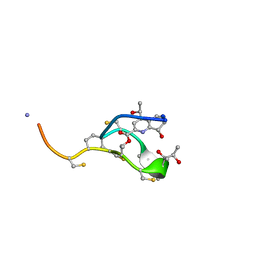 | | Thiostrepton | | Descriptor: | Thiostrepton | | Authors: | Jonker, H.R.A, Baumann, S, Wolf, A, Schoof, S, Hiller, F, Schulte, K.W, Kirschner, K.N, Schwalbe, H, Arndt, H.-D. | | Deposit date: | 2010-08-27 | | Release date: | 2011-02-02 | | Last modified: | 2013-05-08 | | Method: | SOLUTION NMR | | Cite: | NMR structures of thiostrepton derivatives for characterization of the ribosomal binding site.
Angew.Chem.Int.Ed.Engl., 50, 2011
|
|
2L2X
 
 | | Thiostrepton, oxidized at CA-CB bond of residue 9 | | Descriptor: | Thiostrepton | | Authors: | Jonker, H.R.A, Baumann, S, Wolf, A, Schoof, S, Hiller, F, Schulte, K.W, Kirschner, K.N, Schwalbe, H, Arndt, H.-D. | | Deposit date: | 2010-08-27 | | Release date: | 2011-02-02 | | Last modified: | 2013-06-26 | | Method: | SOLUTION NMR | | Cite: | NMR structures of thiostrepton derivatives for characterization of the ribosomal binding site.
Angew.Chem.Int.Ed.Engl., 50, 2011
|
|
2L2Y
 
 | | Thiostrepton, epimer form of residue 9 | | Descriptor: | Thiostrepton | | Authors: | Jonker, H.R.A, Baumann, S, Wolf, A, Schoof, S, Hiller, F, Schulte, K.W, Kirschner, K.N, Schwalbe, H, Arndt, H.-D. | | Deposit date: | 2010-08-27 | | Release date: | 2011-02-02 | | Last modified: | 2013-06-26 | | Method: | SOLUTION NMR | | Cite: | NMR structures of thiostrepton derivatives for characterization of the ribosomal binding site.
Angew.Chem.Int.Ed.Engl., 50, 2011
|
|
3H0B
 
 | | Discovery of aminoheterocycles as a novel beta-secretase inhibitor class | | Descriptor: | 4-[(1S)-1-(3-fluoro-4-methoxyphenyl)-2-(2-methoxy-5-nitrophenyl)ethyl]-1H-imidazol-2-amine, Beta-secretase 1 | | Authors: | Allison, T.J. | | Deposit date: | 2009-04-08 | | Release date: | 2009-07-07 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Discovery of aminoheterocycles as a novel beta-secretase inhibitor class: pH dependence on binding activity part 1.
Bioorg.Med.Chem.Lett., 19, 2009
|
|
3UFL
 
 | | Discovery of Pyrrolidine-based b-Secretase Inhibitors: Lead Advancement through Conformational Design for Maintenance of Ligand Binding Efficiency | | Descriptor: | (1R,4'S)-3,4-dihydro-2H-spiro[naphthalene-1,3'-pyrrolidin]-4'-yl[(2S,4R)-2,4-diphenylpiperidin-1-yl]methanone, Beta-secretase 1, GLYCEROL, ... | | Authors: | Allison, T, Munshi, S, Soisson, S.M. | | Deposit date: | 2011-11-01 | | Release date: | 2012-01-18 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Discovery of pyrrolidine-based beta-secretase inhibitors: Lead advancement through conformational design for maintenance of ligand binding efficiency.
Bioorg.Med.Chem.Lett., 22, 2012
|
|
8CGC
 
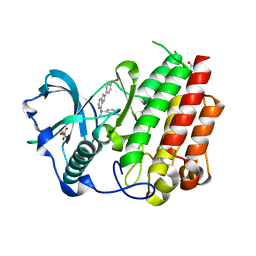 | | Structure of CSF1R in complex with a pyrollopyrimidine (compound 23) | | Descriptor: | (2S)-2-hydroxybutanedioic acid, GLYCEROL, Macrophage colony-stimulating factor 1 receptor, ... | | Authors: | Aarhus, T.I, Bjornstad, F, Wolowczyk, C, Larsen, K.U, Rognstad, L, Leithaug, T, Unger, A, Habenberger, P, Wolff, A, Bjorkoy, G, Pridans, C, Eickhoff, J, Klebl, B, Hoff, B.H, Sundby, E. | | Deposit date: | 2023-02-03 | | Release date: | 2023-05-24 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.925 Å) | | Cite: | Synthesis and Development of Highly Selective Pyrrolo[2,3- d ]pyrimidine CSF1R Inhibitors Targeting the Autoinhibited Form.
J.Med.Chem., 66, 2023
|
|
6EW9
 
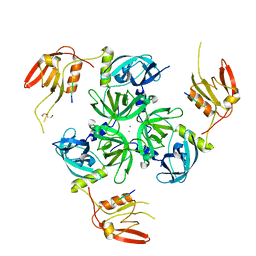 | | CRYSTAL STRUCTURE OF DEGS STRESS SENSOR PROTEASE IN COMPLEX WITH ACTIVATING DNRLGLVYQF PEPTIDE | | Descriptor: | CALCIUM ION, DI(HYDROXYETHYL)ETHER, DNRLGLVYQF PEPTIDE, ... | | Authors: | Vetter, I.R, Porfetye, A.T, Stege, P. | | Deposit date: | 2017-11-03 | | Release date: | 2018-04-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Identification of Noncatalytic Lysine Residues from Allosteric Circuits via Covalent Probes.
ACS Chem. Biol., 13, 2018
|
|
1SI9
 
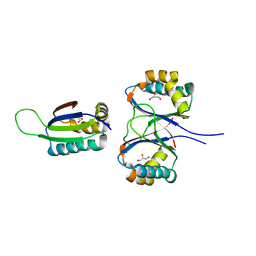 | | Boiling stable protein isolated from Populus tremula | | Descriptor: | GLYCEROL, stable protein 1 | | Authors: | Almog, O, Gonzales, A, Shoseyov, O, Dgany, O, Sofer, O, Wolf, S.G. | | Deposit date: | 2004-02-29 | | Release date: | 2004-09-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | The Structural Basis of the Thermostability of SP1, a Novel Plant (Populus tremula) Boiling Stable Protein.
J.Biol.Chem., 279, 2004
|
|
1TR0
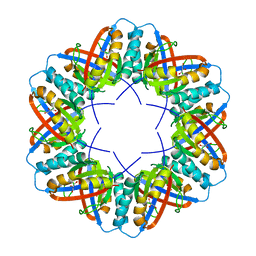 
 | | Crystal Structure of a boiling stable protein SP1 | | Descriptor: | GLYCEROL, stable protein 1 | | Authors: | Almog, O, Gonzalez, A, Sofer, O, Dgany, O, Shoseyov, O. | | Deposit date: | 2004-06-18 | | Release date: | 2004-09-21 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The structural basis of the thermostability of SP1, a novel plant (Populus tremula) boiling stable protein
J.Biol.Chem., 279, 2004
|
|
5JCZ
 
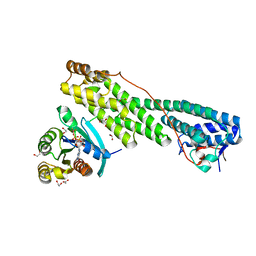 | | Rab11 bound to MyoVa-GTD | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, BERYLLIUM TRIFLUORIDE ION, ... | | Authors: | Pylypenko, O, Attanda, W, Gauquelin, C, Malherbes, G, Houdusse, A. | | Deposit date: | 2016-04-15 | | Release date: | 2016-09-28 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.056 Å) | | Cite: | Coordinated recruitment of Spir actin nucleators and myosin V motors to Rab11 vesicle membranes.
Elife, 5, 2016
|
|
5JCY
 
 | | Spir2-GTBM bound to MyoVa-GTD | | Descriptor: | Protein spire homolog 2, Unconventional myosin-Va | | Authors: | Pylypenko, O, Malherbes, G, Welz, T, Kerkhoff, E, Houdusse, A. | | Deposit date: | 2016-04-15 | | Release date: | 2016-09-28 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Coordinated recruitment of Spir actin nucleators and myosin V motors to Rab11 vesicle membranes.
Elife, 5, 2016
|
|
7OBH
 
 | |
7OBX
 
 | |
7OBY
 
 | |
7OBG
 
 | |
7OBS
 
 | |
7OBD
 
 | |
7OB8
 
 | |
7OBT
 
 | |
7OBK
 
 | |
7OB5
 
 | |
