4F7I
 
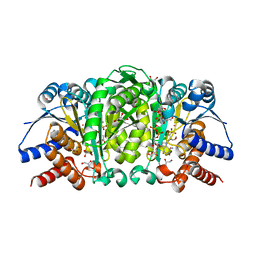 | | Structure of Isopropylmalate dehydrogenase from Thermus thermophilus in complex with IPM, Mn and NADH | | Descriptor: | 3-ISOPROPYLMALIC ACID, 3-isopropylmalate dehydrogenase, 3[N-MORPHOLINO]PROPANE SULFONIC ACID, ... | | Authors: | Pallo, A, Graczer, E, Zavodszky, P, Weiss, M.S, Vas, M. | | Deposit date: | 2012-05-16 | | Release date: | 2012-06-13 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and energetic basis of isopropylmalate dehydrogenase enzyme catalysis.
Febs J., 281, 2014
|
|
4WUO
 
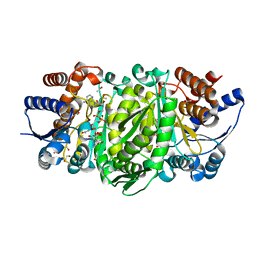 | | Structure of the E270A Mutant Isopropylmalate dehydrogenase from Thermus thermophilus in complex with IPM, Mn and NADH | | Descriptor: | 3-ISOPROPYLMALIC ACID, 3-isopropylmalate dehydrogenase, ETHANOL, ... | | Authors: | Pallo, A, Graczer, E, Olah, J, Szimler, T, Konarev, P.V, Svergun, D.I, Merli, A, Zavodszky, P, Vas, M, Weiss, M.S. | | Deposit date: | 2014-11-03 | | Release date: | 2014-11-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Glutamate 270 plays an essential role in K(+)-activation and domain closure of Thermus thermophilus isopropylmalate dehydrogenase.
Febs Lett., 589, 2015
|
|
5C04
 
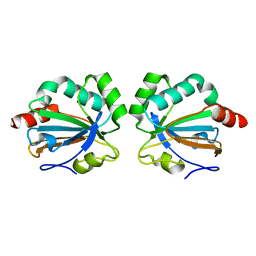 | |
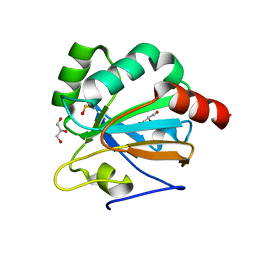
4X1U
 
 | |
4X0X
 
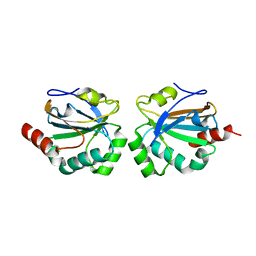 | |
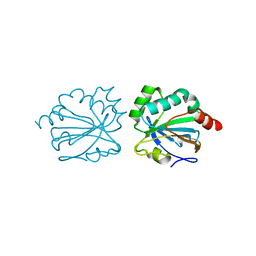
4XIH
 
 | |
5LOL
 
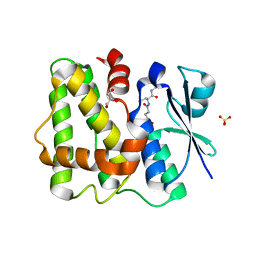 | | Glutathione-bound Dehydroascorbate Reductase 2 of Arabidopsis thaliana | | Descriptor: | GLUTATHIONE, GLYCEROL, Glutathione S-transferase DHAR2, ... | | Authors: | Young, D.R, Pallo, A, Bodra, N, Messens, J. | | Deposit date: | 2016-08-09 | | Release date: | 2017-06-07 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Arabidopsis thaliana dehydroascorbate reductase 2: Conformational flexibility during catalysis.
Sci Rep, 7, 2017
|
|
2QR5
 
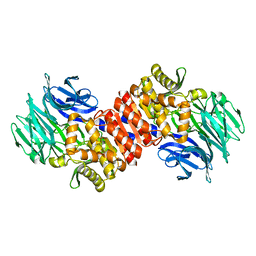 | | Aeropyrum pernix acylaminoacyl peptidase, H367A mutant | | Descriptor: | Acylamino-acid-releasing enzyme | | Authors: | Harmat, V, Pallo, A, Kiss, A.L, Polgar, L. | | Deposit date: | 2007-07-27 | | Release date: | 2008-05-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural and kinetic contributions of the oxyanion binding site to the catalytic activity of acylaminoacyl peptidase
J.Struct.Biol., 162, 2008
|
|
2QY0
 
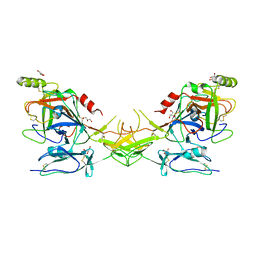 | | Active dimeric structure of the catalytic domain of C1r reveals enzyme-product like contacts | | Descriptor: | Complement C1r subcomponent, GLYCEROL | | Authors: | Kardos, J, Harmat, V, Pallo, A, Barabas, O, Szilagyi, K, Graf, L, Naray-Szabo, G, Goto, Y, Zavodszky, P, Gal, P. | | Deposit date: | 2007-08-13 | | Release date: | 2008-02-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Revisiting the mechanism of the autoactivation of the complement protease C1r in the C1 complex: Structure of the active catalytic region of C1r.
Mol.Immunol., 45, 2008
|
|
3O4J
 
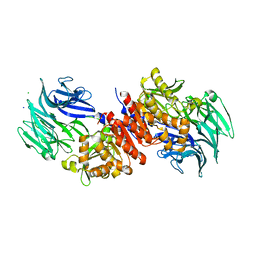 | | Structure and Catalysis of Acylaminoacyl Peptidase | | Descriptor: | Acylamino-acid-releasing enzyme, CHLORIDE ION, GLYCEROL, ... | | Authors: | Harmat, V, Domokos, K, Menyhard, D.K, Pallo, A, Szeltner, Z, Szamosi, I, Beke-Somfai, T, Naray-Szabo, G, Polgar, L. | | Deposit date: | 2010-07-27 | | Release date: | 2010-11-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure and Catalysis of Acylaminoacyl Peptidase: CLOSED AND OPEN SUBUNITS OF A DIMER OLIGOPEPTIDASE.
J.Biol.Chem., 286, 2011
|
|
3O4I
 
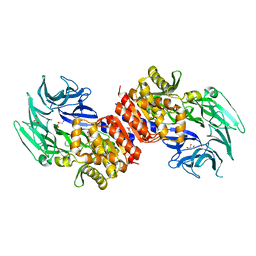 | | Structure and Catalysis of Acylaminoacyl Peptidase | | Descriptor: | Acylamino-acid-releasing enzyme, CHLORIDE ION, GLYCEROL | | Authors: | Harmat, V, Domokos, K, Menyhard, D.K, Pallo, A, Szeltner, Z, Szamosi, I, Beke-Somfai, T, Naray-Szabo, G, Polgar, L. | | Deposit date: | 2010-07-27 | | Release date: | 2010-11-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure and Catalysis of Acylaminoacyl Peptidase: CLOSED AND OPEN SUBUNITS OF A DIMER OLIGOPEPTIDASE.
J.Biol.Chem., 286, 2011
|
|
3O4G
 
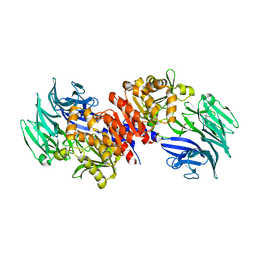 | | Structure and Catalysis of Acylaminoacyl Peptidase | | Descriptor: | Acylamino-acid-releasing enzyme, GLYCEROL | | Authors: | Harmat, V, Domokos, K, Menyhard, D.K, Pallo, A, Szeltner, Z, Szamosi, I, Beke-Somfai, T, Naray-Szabo, G, Polgar, L. | | Deposit date: | 2010-07-27 | | Release date: | 2010-11-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure and Catalysis of Acylaminoacyl Peptidase: CLOSED AND OPEN SUBUNITS OF A DIMER OLIGOPEPTIDASE.
J.Biol.Chem., 286, 2011
|
|
3O4H
 
 | | Structure and Catalysis of Acylaminoacyl Peptidase | | Descriptor: | Acylamino-acid-releasing enzyme, GLYCEROL, SODIUM ION | | Authors: | Harmat, V, Domokos, K, Menyhard, D.K, Pallo, A, Szeltner, Z, Szamosi, I, Beke-Somfai, T, Naray-Szabo, G, Polgar, L. | | Deposit date: | 2010-07-27 | | Release date: | 2010-11-17 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structure and Catalysis of Acylaminoacyl Peptidase: CLOSED AND OPEN SUBUNITS OF A DIMER OLIGOPEPTIDASE.
J.Biol.Chem., 286, 2011
|
|
