5X9K
 
 | |
5X9J
 
 | |
5WQH
 
 | | Structure of fungal meroterpenoid isomerase Trt14 complexed with substrate analog and endo-terretonin D | | Descriptor: | CALCIUM ION, Isomerase trt14, methyl (2S,4aR,4bS,5S,6aS,10aS,10bS,12aS)-2,4b,7,7,10a,12,12a-heptamethyl-5-oxidanyl-1,4,6,8-tetrakis(oxidanylidene)-4a,5,6a,9,10,10b-hexahydronaphtho[1,2-h]isochromene-2-carboxylate, ... | | Authors: | Mori, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-26 | | Release date: | 2017-07-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.102 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
5WQG
 
 | | Structure of fungal meroterpenoid isomerase Trt14 complexed with terretonin D | | Descriptor: | CALCIUM ION, Isomerase trt14, methyl (2S,4aR,4bS,5S,6aS,10aS,10bS,12aS)-2,4b,7,7,10a,12a-hexamethyl-12-methylidene-5-oxidanyl-1,4,6,8-tetrakis(oxidanylidene) -5,6a,9,10,10b,11-hexahydro-4aH-naphtho[1,2-h]isochromene-2-carboxylate | | Authors: | Mori, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-26 | | Release date: | 2017-07-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
5WQF
 
 | | Structure of fungal meroterpenoid isomerase Trt14 | | Descriptor: | CALCIUM ION, Isomerase trt14, N-PROPANOL | | Authors: | Mori, T, Iwabuchi, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-26 | | Release date: | 2017-07-26 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
5WQI
 
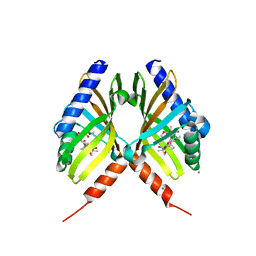 | | Structure of fungal meroterpenoid isomerase Trt14 complexed with hydrolyzed product | | Descriptor: | (1R,2S,4aS,4bS,8aS,10S,10aS)-2-[(2S)-3-methoxy-2-methyl-2-oxidanyl-3-oxidanylidene-propanoyl]-2,3,4b,8,8,10a-hexamethyl-10-oxidanyl-7,9-bis(oxidanylidene)-1,4a,5,6,8a,10-hexahydrophenanthrene-1-carboxylic acid, CALCIUM ION, Isomerase trt14 | | Authors: | Mori, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-27 | | Release date: | 2017-07-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.899 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
1WPK
 
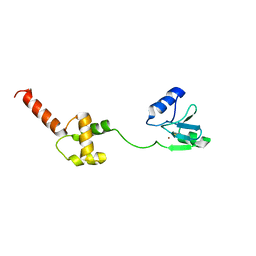 | | Methylated Form of N-terminal Transcriptional Regulator Domain of Escherichia Coli Ada Protein | | Descriptor: | ADA regulatory protein, ZINC ION | | Authors: | Takinowaki, H, Matsuda, Y, Yoshida, T, Kobayashi, Y, Ohkubo, T. | | Deposit date: | 2004-09-07 | | Release date: | 2005-09-13 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | The solution structure of the methylated form of the N-terminal 16-kDa domain of Escherichia coli Ada protein
Protein Sci., 15, 2006
|
|
2D35
 
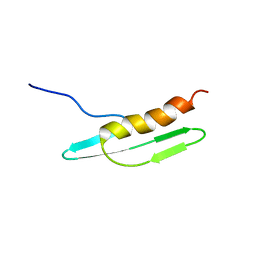 | | Solution structure of Cell Division Reactivation Factor, CedA | | Descriptor: | Cell division activator cedA | | Authors: | Abe, Y, Watanabe, N, Matsuda, Y, Yoshida, Y, Katayama, T, Ueda, T. | | Deposit date: | 2005-09-26 | | Release date: | 2006-12-12 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural Analysis and Molecular Interaction of Cell Division Reactivation Factor, CedA from Escherichia coli
To be Published
|
|
7DEM
 
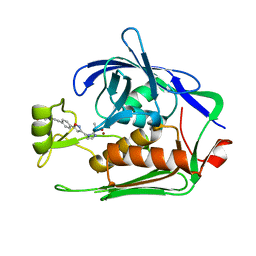 | | Crystal structure of P.aeruginosa LpxC in complex with inhibitor | | Descriptor: | 5-[4-[3-[[2-[(1~{S})-1-oxidanylethyl]imidazol-1-yl]methyl]-1,2-oxazol-5-yl]phenyl]pent-4-yn-1-ol, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ZINC ION | | Authors: | Mima, M, Ushiyama, F, Matsuda, Y. | | Deposit date: | 2020-11-04 | | Release date: | 2021-01-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Lead optimization of 2-hydroxymethyl imidazoles as non-hydroxamate LpxC inhibitors: Discovery of TP0586532.
Bioorg.Med.Chem., 30, 2020
|
|
6LZH
 
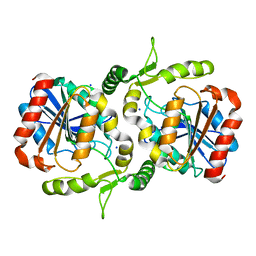 | | Crystal structure of Alpha/beta hydrolase GrgF from Penicillium sp. sh18 | | Descriptor: | GrgF, SODIUM ION | | Authors: | Wang, H, Yu, J, Wang, W.G, Matsuda, Y, Yao, M. | | Deposit date: | 2020-02-19 | | Release date: | 2020-06-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular Basis for the Biosynthesis of an Unusual Chain-Fused Polyketide, Gregatin A.
J.Am.Chem.Soc., 142, 2020
|
|
4XYD
 
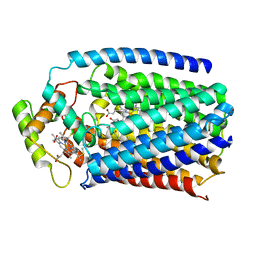 | | Nitric oxide reductase from Roseobacter denitrificans (RdNOR) | | Descriptor: | CALCIUM ION, COPPER (II) ION, FE (III) ION, ... | | Authors: | Crow, A. | | Deposit date: | 2015-02-02 | | Release date: | 2016-05-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structure of the Membrane-intrinsic Nitric Oxide Reductase from Roseobacter denitrificans.
Biochemistry, 55, 2016
|
|
8X0V
 
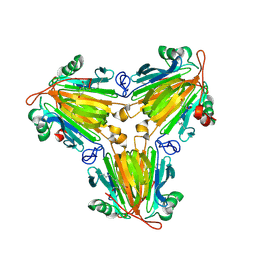 | |
8X0U
 
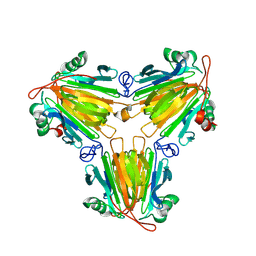 | |
7F0Y
 
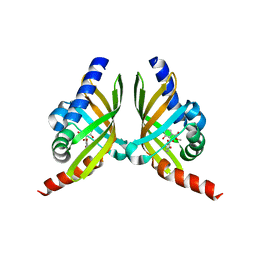 | |
7F13
 
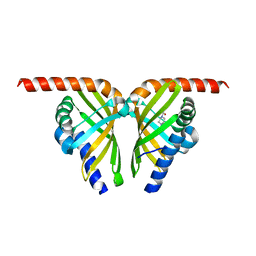 | | Crystal structure of isomerase Dcr3 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Dcr3 | | Authors: | Yang, J, Mori, T, Abe, I. | | Deposit date: | 2021-06-07 | | Release date: | 2022-04-20 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural Basis for Isomerization Reactions in Fungal Tetrahydroxanthone Biosynthesis and Diversification.
Angew.Chem.Int.Ed.Engl., 60, 2021
|
|
7F11
 
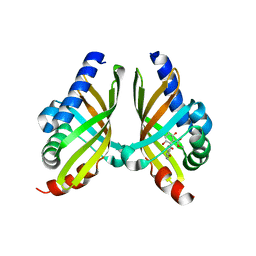 | | Crystal structure of NsrQ M128I in complex with substrate analogue 7 | | Descriptor: | NsrQ, methyl 2-[2,6-bis(oxidanyl)phenyl]carbonyl-5-methyl-3,6-bis(oxidanyl)benzoate | | Authors: | Yang, J, Mori, T, Abe, I. | | Deposit date: | 2021-06-07 | | Release date: | 2022-04-20 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural Basis for Isomerization Reactions in Fungal Tetrahydroxanthone Biosynthesis and Diversification.
Angew.Chem.Int.Ed.Engl., 60, 2021
|
|
7F0Z
 
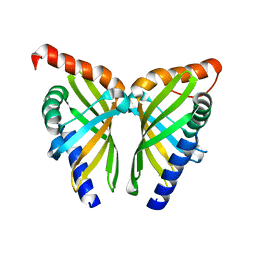 | | Crystal structure of NsrQ W31A | | Descriptor: | NsrQ | | Authors: | Yang, J, Mori, T, Abe, I. | | Deposit date: | 2021-06-07 | | Release date: | 2022-04-20 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for Isomerization Reactions in Fungal Tetrahydroxanthone Biosynthesis and Diversification.
Angew.Chem.Int.Ed.Engl., 60, 2021
|
|
7F0O
 
 | |
7F14
 
 | |
7F10
 
 | |
2E0G
 
 | |
7CID
 
 | | Crystal structure of P.aeruginosa LpxC in complex with inhibitor | | Descriptor: | 1-[3-(4-chlorophenyl)propyl]imidazole, DIMETHYL SULFOXIDE, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ... | | Authors: | Baker, L.M, Mima, M, Surgenor, A, Robertson, A. | | Deposit date: | 2020-07-07 | | Release date: | 2020-12-02 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Fragment-Based Discovery of Novel Non-Hydroxamate LpxC Inhibitors with Antibacterial Activity.
J.Med.Chem., 63, 2020
|
|
7CI4
 
 | | Crystal structure of P.aeruginosa LpxC in complex with inhibitor | | Descriptor: | (2R)-2-azanyl-4-methylsulfonyl-N-[3-(trifluoromethyloxy)phenyl]butanamide, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Mima, M, Baker, L.M, Surgenor, A, Robertson, A. | | Deposit date: | 2020-07-07 | | Release date: | 2020-12-02 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Fragment-Based Discovery of Novel Non-Hydroxamate LpxC Inhibitors with Antibacterial Activity.
J.Med.Chem., 63, 2020
|
|
7CIC
 
 | | Crystal structure of P.aeruginosa LpxC in complex with inhibitor | | Descriptor: | 2-azanyl-N-[3-(trifluoromethyloxy)phenyl]ethanamide, UDP-3-O-acyl-N-acetylglucosamine deacetylase, ZINC ION | | Authors: | Baker, L.M, Mima, M, Surgenor, A, Robertson, A. | | Deposit date: | 2020-07-07 | | Release date: | 2020-12-02 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Fragment-Based Discovery of Novel Non-Hydroxamate LpxC Inhibitors with Antibacterial Activity.
J.Med.Chem., 63, 2020
|
|
7CI9
 
 | | Crystal structure of P.aeruginosa LpxC in complex with inhibitor | | Descriptor: | 1,2-ETHANEDIOL, 2-azanyl-2-[[4-[2-[3-[[2-[(1S)-1-oxidanylethyl]imidazol-1-yl]methyl]-1,2-oxazol-5-yl]ethynyl]phenoxy]methyl]propane-1,3-diol, PHOSPHATE ION, ... | | Authors: | Mima, M, Baker, L.M, Surgenor, A, Robertson, A. | | Deposit date: | 2020-07-07 | | Release date: | 2020-12-02 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Fragment-Based Discovery of Novel Non-Hydroxamate LpxC Inhibitors with Antibacterial Activity.
J.Med.Chem., 63, 2020
|
|
