1IC2
 
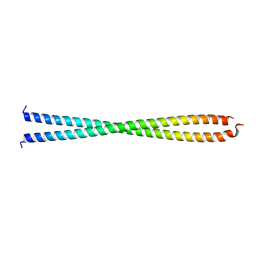 | | DECIPHERING THE DESIGN OF THE TROPOMYOSIN MOLECULE | | Descriptor: | TROPOMYOSIN ALPHA CHAIN, SKELETAL MUSCLE | | Authors: | Brown, J.H, Kim, K.-H, Jun, G, Greenfield, N.J, Dominguez, R, Volkmann, N, Hitchcock-DeGregori, S.E, Cohen, C. | | Deposit date: | 2001-03-29 | | Release date: | 2001-07-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Deciphering the design of the tropomyosin molecule
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1DEQ
 
 | | THE CRYSTAL STRUCTURE OF MODIFIED BOVINE FIBRINOGEN (AT ~4 ANGSTROM RESOLUTION) | | Descriptor: | FIBRINOGEN, FIBRINOGEN (ALPHA CHAIN), FIBRINOGEN (BETA CHAIN), ... | | Authors: | Brown, J.H, Volkmann, N, Jun, G, Henschen-Edman, A.H, Cohen, C. | | Deposit date: | 1999-11-15 | | Release date: | 2000-02-02 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | The crystal structure of modified bovine fibrinogen.
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
1AJ1
 
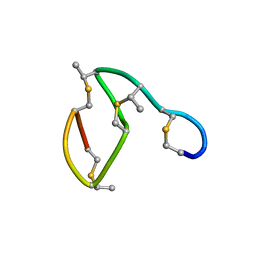 | | NMR STRUCTURE OF THE LANTIBIOTIC ACTAGARDINE | | Descriptor: | LANTIBIOTIC ACTAGARDINE | | Authors: | Zimmermann, N, Jung, G. | | Deposit date: | 1997-05-14 | | Release date: | 1997-10-15 | | Last modified: | 2024-07-10 | | Method: | SOLUTION NMR | | Cite: | The three-dimensional solution structure of the lantibiotic murein-biosynthesis-inhibitor actagardine determined by NMR.
Eur.J.Biochem., 246, 1997
|
|
2P1B
 
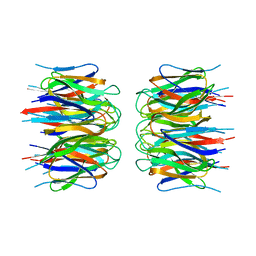 | | Crystal structure of human nucleophosmin-core | | Descriptor: | Nucleophosmin | | Authors: | Lee, H.H, Kim, H.S, Kang, J.Y, Lee, B.I, Ha, J.Y, Yoon, H.J, Lim, S.O, Jung, G, Suh, S.W. | | Deposit date: | 2007-03-03 | | Release date: | 2007-03-27 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Crystal structure of human nucleophosmin-core reveals plasticity of the pentamer-pentamer interface
Proteins, 69, 2007
|
|
4P1Q
 
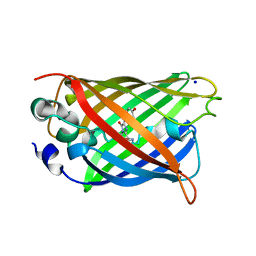 | | GREEN FLUORESCENT PROTEIN E222H VARIANT | | Descriptor: | Green fluorescent protein, SODIUM ION | | Authors: | Klein, M, Carius, Y, Auerbach, D, Franz, S, Jung, G, Lancaster, C.R.D. | | Deposit date: | 2014-02-27 | | Release date: | 2014-07-16 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Replacement of Highly Conserved E222 by the Photostable Non-photoconvertible Histidine in GFP.
Chembiochem, 15, 2014
|
|
1OXE
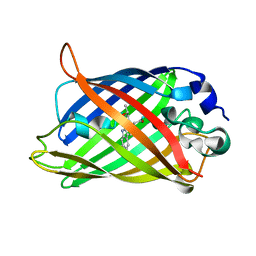 
 | | Expansion of the Genetic Code Enables Design of a Novel "Gold" Class of Green Fluorescent Proteins | | Descriptor: | cyan fluorescent protein cfp | | Authors: | Hyun Bae, J, Rubini, M, Jung, G, Wiegand, G, Seifert, M.H, Azim, M.K, Kim, J.S, Zumbusch, A, Holak, T.A, Moroder, L, Huber, R, Budisa, N. | | Deposit date: | 2003-04-02 | | Release date: | 2003-12-02 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Expansion of the Genetic Code Enables Design of a Novel "Gold" Class of Green Fluorescent Proteins
J.Mol.Biol., 328, 2003
|
|
1OXF
 
 | | Expansion of the Genetic Code Enables Design of a Novel "Gold" Class of Green Fluorescent Proteins | | Descriptor: | cyan fluorescent protein cfp | | Authors: | Hyun Bae, J, Rubini, M, Jung, G, Wiegand, G, Seifert, M.H, Azim, M.K, Kim, J.S, Zumbusch, A, Holak, T.A, Moroder, L, Huber, R, Budisa, N. | | Deposit date: | 2003-04-02 | | Release date: | 2003-12-02 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Expansion of the Genetic Code Enables Design of a Novel "Gold" Class of Green Fluorescent Proteins
J.Mol.Biol., 328, 2003
|
|
1OXD
 
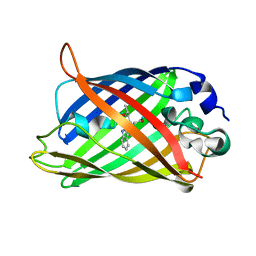 | | Expansion of the Genetic Code Enables Design of a Novel "Gold" Class of Green Fluorescent Proteins | | Descriptor: | cyan fluorescent protein cfp | | Authors: | Hyun Bae, J, Rubini, M, Jung, G, Wiegand, G, Seifert, M.H, Azim, M.K, Kim, J.S, Zumbusch, A, Holak, T.A, Moroder, L, Huber, R, Budisa, N. | | Deposit date: | 2003-04-02 | | Release date: | 2003-12-02 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Expansion of the Genetic Code Enables Design of a Novel "Gold" Class of Green Fluorescent Proteins
J.Mol.Biol., 328, 2003
|
|
1G6E
 
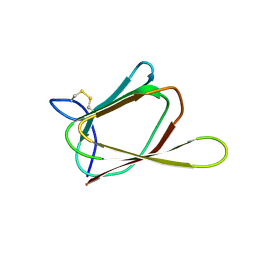 | | ANTIFUNGAL PROTEIN FROM STREPTOMYCES TENDAE TU901, 30-CONFORMERS ENSEMBLE | | Descriptor: | ANTIFUNGAL PROTEIN | | Authors: | Campos-Olivas, R, Bormann, C, Hoerr, I, Jung, G, Gronenborn, A.M. | | Deposit date: | 2000-11-04 | | Release date: | 2001-03-28 | | Last modified: | 2022-12-21 | | Method: | SOLUTION NMR | | Cite: | Solution structure, backbone dynamics and chitin binding of the anti-fungal protein from Streptomyces tendae TU901.
J.Mol.Biol., 308, 2001
|
|
1GH5
 
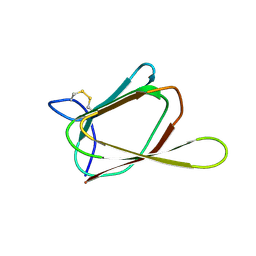 | | ANTIFUNGAL PROTEIN FROM STREPTOMYCES TENDAE TU901, NMR AVERAGE STRUCTURE | | Descriptor: | ANTIFUNGAL PROTEIN | | Authors: | Campos-Olivas, R, Bormann, C, Hoerr, I, Jung, G, Gronenborn, A.M. | | Deposit date: | 2000-11-04 | | Release date: | 2001-03-28 | | Last modified: | 2022-12-21 | | Method: | SOLUTION NMR | | Cite: | Solution structure, backbone dynamics and chitin binding of the anti-fungal protein from Streptomyces tendae TU901.
J.Mol.Biol., 308, 2001
|
|
3PDO
 
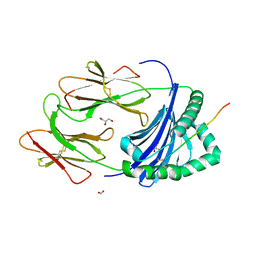 | | Crystal Structure of HLA-DR1 with CLIP102-120 | | Descriptor: | FORMIC ACID, GLYCEROL, HLA class II histocompatibility antigen gamma chain, ... | | Authors: | Gunther, S, Schlundt, A, Sticht, J, Roske, Y, Heinemann, U, Wiesmuller, K.-H, Jung, G, Falk, K, Rotzschke, O, Freund, C. | | Deposit date: | 2010-10-23 | | Release date: | 2010-12-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Bidirectional binding of invariant chain peptides to an MHC class II molecule.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3PGC
 
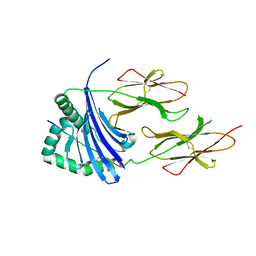 | | Crystal Structure of HLA-DR1 with CLIP106-120, flipped peptide orientation | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, HLA class II histocompatibility antigen gamma chain, ... | | Authors: | Gunther, S, Schlundt, A, Sticht, J, Roske, Y, Heinemann, U, Wiesmuller, K.-H, Jung, G, Falk, K, Rotzschke, O, Freund, C. | | Deposit date: | 2010-11-01 | | Release date: | 2010-12-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.66 Å) | | Cite: | Bidirectional binding of invariant chain peptides to an MHC class II molecule.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3PGD
 
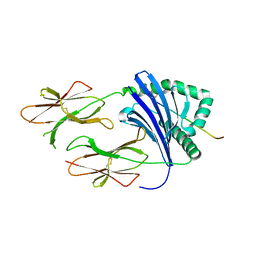 | | Crystal Structure of HLA-DR1 with CLIP106-120, canonical peptide orientation | | Descriptor: | HLA class II histocompatibility antigen gamma chain, HLA class II histocompatibility antigen, DR alpha chain, ... | | Authors: | Gunther, S, Schlundt, A, Sticht, J, Roske, Y, Heinemann, U, Wiesmuller, K.-H, Jung, G, Falk, K, Rotzschke, O, Freund, C. | | Deposit date: | 2010-11-01 | | Release date: | 2010-12-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.72 Å) | | Cite: | Bidirectional binding of invariant chain peptides to an MHC class II molecule.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
6L6Z
 
 | |
6LFG
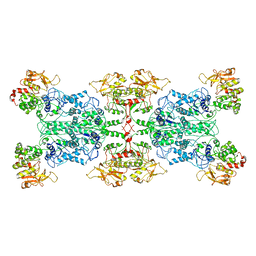 
 | |
1LGR
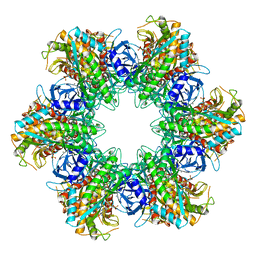 
 | |
1JY2
 
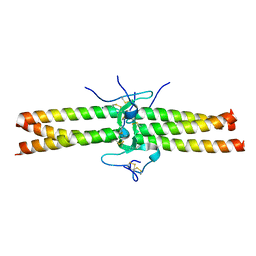 | | Crystal Structure of the Central Region of Bovine Fibrinogen (E5 fragment) at 1.4 Angstroms Resolution | | Descriptor: | FIBRINOGEN ALPHA CHAIN, FIBRINOGEN BETA CHAIN, FIBRINOGEN GAMMA-B CHAIN | | Authors: | Madrazo, J, Brown, J.H, Litvinovich, S, Dominguez, R, Yakovlev, S, Medved, L, Cohen, C. | | Deposit date: | 2001-09-10 | | Release date: | 2001-10-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystal structure of the central region of bovine fibrinogen (E5 fragment) at 1.4-A resolution.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1JY3
 
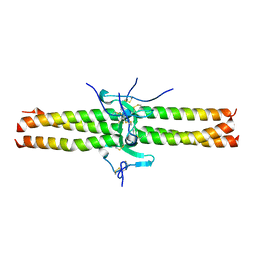 | | Crystal Structure of the Central Region of Bovine Fibrinogen (E5 Fragment) at 1.4 Angstroms Resolution | | Descriptor: | FIBRINOGEN ALPHA CHAIN, FIBRINOGEN BETA CHAIN, FIBRINOGEN GAMMA-B CHAIN | | Authors: | Madrazo, J, Brown, J.H, Litvinovich, S, Dominguez, R, Yakovlev, S, Medved, L, Cohen, C. | | Deposit date: | 2001-09-10 | | Release date: | 2001-10-17 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of the central region of bovine fibrinogen (E5 fragment) at 1.4-A resolution.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
4PFE
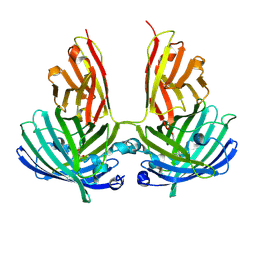 
 | | Crystal structure of vsfGFP-0 | | Descriptor: | Green fluorescent protein | | Authors: | Jauch, R, Chen, S.L. | | Deposit date: | 2014-04-29 | | Release date: | 2015-06-24 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.603 Å) | | Cite: | Rational Structure-Based Design of Bright GFP-Based Complexes with Tunable Dimerization.
Angew.Chem.Int.Ed.Engl., 54, 2015
|
|
3LO2
 
 | |
3LO1
 
 | |
3LOE
 
 | |
3LO4
 
 | |
3LO6
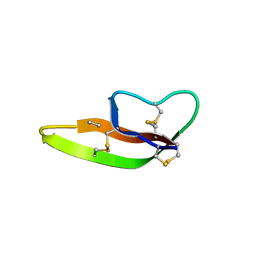 
 | |
3LO9
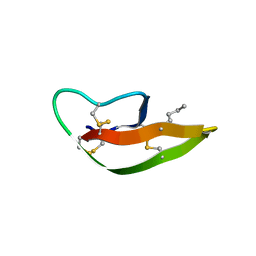 
 | |
