9W4S
 
 | | Cryo-EM structure of hTUT4_mini:hLin28A:pre-let-7g miRNA_UUU, conformation 2 | | Descriptor: | Protein lin-28 homolog A, Terminal uridylyltransferase 4, ZINC ION, ... | | Authors: | Han, X, Yamashita, S, Tomita, K. | | Deposit date: | 2025-08-01 | | Release date: | 2025-12-03 | | Last modified: | 2026-01-28 | | Method: | ELECTRON MICROSCOPY (3.82 Å) | | Cite: | Mechanistic insights into Lin28-dependent oligo-uridylylation of pre-let-7 by TUT4.
Nucleic Acids Res., 54, 2026
|
|
9W4R
 
 | | Cryo-EM structure of hTUT4_mini:hLin28A:pre-let-7g miRNA_UUU, conformation 1 | | Descriptor: | Protein lin-28 homolog A, Terminal uridylyltransferase 4, ZINC ION, ... | | Authors: | Han, X, Yamashita, S, Tomita, K. | | Deposit date: | 2025-08-01 | | Release date: | 2025-12-03 | | Last modified: | 2026-01-28 | | Method: | ELECTRON MICROSCOPY (3.78 Å) | | Cite: | Mechanistic insights into Lin28-dependent oligo-uridylylation of pre-let-7 by TUT4.
Nucleic Acids Res., 54, 2026
|
|
9W6W
 
 | | 5-CT-bound serotonin 5A (5-HT5A) receptor-Gi protein complex | | Descriptor: | 3-(2-azanylethyl)-1H-indole-5-carboxamide, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, ... | | Authors: | Zhang, X, Xie, L, Chen, P, Yu, J, Tian, X, Wei, J, Shao, Z, Yan, W, Xu, Z. | | Deposit date: | 2025-08-05 | | Release date: | 2025-11-26 | | Method: | ELECTRON MICROSCOPY (3.13 Å) | | Cite: | Structural and Functional Basis for Biased Agonism at the 5-hydroxytryptamine 5A Receptor.
Mol Biomed, 6, 2025
|
|
9UAZ
 
 | |
9UCP
 
 | | cryo-EM structure of M1 muscarinic acetylcholine receptor-alpha5 helix of G11 protein complex bound to iperoxo and nanobody Nb1B4 | | Descriptor: | 4-(4,5-dihydro-1,2-oxazol-3-yloxy)-N,N,N-trimethylbut-2-yn-1-aminium, Guanine nucleotide-binding protein subunit alpha-11, M1 muscarinic acetylcholine receptor, ... | | Authors: | Zhang, X, Gao, K, Liu, X. | | Deposit date: | 2025-04-05 | | Release date: | 2025-10-29 | | Method: | ELECTRON MICROSCOPY (2.88 Å) | | Cite: | cryo-EM structure of M1 muscarinic acetylcholine receptor-alpha5 helix of G11 protein complex bound to iperoxo and nanobody Nb1B4
To Be Published
|
|
9UAP
 
 | |
9WFH
 
 | | Carbohydrate-binding module 32 of LnbB from Bifidobacterium bifidum, ligand free form, multiple small-wedge data set | | Descriptor: | CALCIUM ION, Lacto-N-biosidase | | Authors: | Zhang, X, Kashima, T, Fushinobu, S. | | Deposit date: | 2025-08-21 | | Release date: | 2025-11-19 | | Last modified: | 2026-03-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insights into lacto-N-biose I recognition by a family 32 carbohydrate-binding module from Bifidobacterium bifidum.
Febs Lett., 600, 2026
|
|
9VAK
 
 | | Carbohydrate-binding module 32 of LnbB from Bifidobacterium bifidum, LNB complex | | Descriptor: | CALCIUM ION, Lacto-N-biosidase, beta-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Zhang, X, Kashima, T, Fushinobu, S. | | Deposit date: | 2025-06-03 | | Release date: | 2025-08-13 | | Last modified: | 2026-03-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insights into lacto-N-biose I recognition by a family 32 carbohydrate-binding module from Bifidobacterium bifidum.
Febs Lett., 600, 2026
|
|
9UPC
 
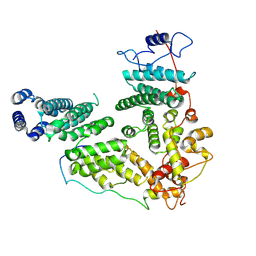 | | Structure of human TREX-2 | | Descriptor: | 26S proteasome complex subunit SEM1, Germinal-center associated nuclear protein, PCI domain-containing protein 2 | | Authors: | Zhang, X, Gong, X, Ge, X. | | Deposit date: | 2025-04-28 | | Release date: | 2026-02-11 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | Structure of human TREX-2
To Be Published
|
|
9UPB
 
 | | Structure of the human TREX-2 bound to UAP56 | | Descriptor: | 26S proteasome complex subunit SEM1, Germinal-center associated nuclear protein, PCI domain-containing protein 2, ... | | Authors: | Zhang, X, Gong, X, Ge, X. | | Deposit date: | 2025-04-28 | | Release date: | 2026-02-18 | | Method: | ELECTRON MICROSCOPY (3.41 Å) | | Cite: | Structure of the human TREX-2 bound to UAP56
To Be Published
|
|
3W28
 
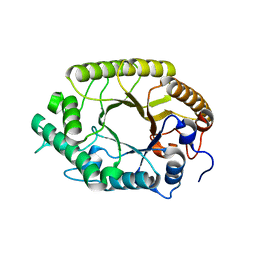 | | The high-resolution crystal structure of TsXylA, intracellular xylanase from /Thermoanaerobacterium saccharolyticum JW/SL-YS485/: the complex of the E251A mutant with xylotriose | | Descriptor: | Glycoside hydrolase family 10, beta-D-xylopyranose-(1-4)-beta-D-xylopyranose-(1-4)-alpha-D-xylopyranose | | Authors: | Han, X, Gao, J, Shang, N, Huang, C.-H, Ko, T.-P, Zhu, Z, Wiegel, J, Shao, W, Guo, R.-T. | | Deposit date: | 2012-11-27 | | Release date: | 2013-04-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Structural and functional analyses of catalytic domain of GH10 xylanase from Thermoanaerobacterium saccharolyticum JW/SL-YS485
Proteins, 81, 2013
|
|
3W24
 
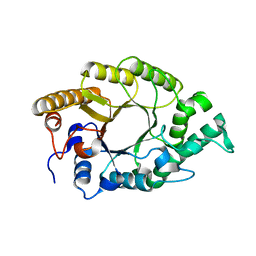 | | The high-resolution crystal structure of TsXylA, intracellular xylanase from Thermoanaerobacterium saccharolyticum JW/SL-YS485 | | Descriptor: | Glycoside hydrolase family 10 | | Authors: | Han, X, Gao, J, Shang, N, Huang, C.-H, Ko, T.-P, Zhu, Z, Wiegel, J, Shao, W, Guo, R.-T. | | Deposit date: | 2012-11-27 | | Release date: | 2013-04-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structural and functional analyses of catalytic domain of GH10 xylanase from Thermoanaerobacterium saccharolyticum JW/SL-YS485
Proteins, 81, 2013
|
|
5V57
 
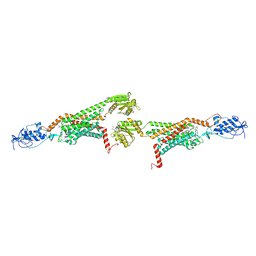 | | 3.0A SYN structure of the multi-domain human smoothened receptor in complex with TC114 | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, FLAVIN MONONUCLEOTIDE, N-methyl-N-[1-[4-(2-methylpyrazol-3-yl)phthalazin-1-yl]piperidin-4-yl]-4-nitro-2-(trifluoromethyl)benzamide, ... | | Authors: | Zhang, X, Zhao, F, Wu, Y, Yang, J, Han, G.W, Zhao, S, Ishchenko, A, Ye, L, Lin, X, Ding, K, Dharmarajan, V, Griffin, P.R, Gati, C, Nelson, G, Hunter, M.S, Hanson, M.A, Cherezov, V, Stevens, R.C, Tan, W, Tao, H, Xu, F. | | Deposit date: | 2017-03-13 | | Release date: | 2017-05-24 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of a multi-domain human smoothened receptor in complex with a super stabilizing ligand.
Nat Commun, 8, 2017
|
|
3W25
 
 | | The high-resolution crystal structure of TsXylA, intracellular xylanase from /Thermoanaerobacterium saccharolyticum JW/SL-YS485/: the complex of the E146A mutant with xylobiose | | Descriptor: | Glycoside hydrolase family 10, beta-D-xylopyranose-(1-4)-beta-D-xylopyranose | | Authors: | Han, X, Gao, J, Shang, N, Huang, C.-H, Ko, T.-P, Zhu, Z, Wiegel, J, Shao, W, Guo, R.-T. | | Deposit date: | 2012-11-27 | | Release date: | 2013-04-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Structural and functional analyses of catalytic domain of GH10 xylanase from Thermoanaerobacterium saccharolyticum JW/SL-YS485
Proteins, 81, 2013
|
|
3W26
 
 | | The high-resolution crystal structure of TsXylA, intracellular xylanase from /Thermoanaerobacterium saccharolyticum JW/SL-YS485/: the complex of the E146A mutant with xylotriose | | Descriptor: | Glycoside hydrolase family 10, beta-D-xylopyranose-(1-4)-beta-D-xylopyranose-(1-4)-beta-D-xylopyranose | | Authors: | Han, X, Gao, J, Shang, N, Huang, C.-H, Ko, T.-P, Zhu, Z, Wiegel, J, Shao, W, Guo, R.-T. | | Deposit date: | 2012-11-27 | | Release date: | 2013-04-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural and functional analyses of catalytic domain of GH10 xylanase from Thermoanaerobacterium saccharolyticum JW/SL-YS485
Proteins, 81, 2013
|
|
5V56
 
 | | 2.9A XFEL structure of the multi-domain human smoothened receptor (with E194M mutation) in complex with TC114 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, FLAVIN MONONUCLEOTIDE, N-methyl-N-[1-[4-(2-methylpyrazol-3-yl)phthalazin-1-yl]piperidin-4-yl]-4-nitro-2-(trifluoromethyl)benzamide, ... | | Authors: | Zhang, X, Zhao, F, Wu, Y, Yang, J, Han, G.W, Zhao, S, Ishchenko, A, Ye, L, Lin, X, Ding, K, Dharmarajan, V, Griffin, P.R, Gati, C, Nelson, G, Hunter, M.S, Hanson, M.A, Cherezov, V, Stevens, R.C, Tan, W, Tao, H, Xu, F. | | Deposit date: | 2017-03-13 | | Release date: | 2017-05-24 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of a multi-domain human smoothened receptor in complex with a super stabilizing ligand.
Nat Commun, 8, 2017
|
|
3W29
 
 | | The high-resolution crystal structure of TsXylA, intracellular xylanase from /Thermoanaerobacterium saccharolyticum JW/SL-YS485/: the complex of the E251A mutant with xylotetraose | | Descriptor: | Glycoside hydrolase family 10, beta-D-xylopyranose-(1-4)-beta-D-xylopyranose-(1-4)-beta-D-xylopyranose-(1-4)-alpha-D-xylopyranose | | Authors: | Han, X, Gao, J, Shang, N, Huang, C.-H, Ko, T.-P, Zhu, Z, Wiegel, J, Shao, W, Guo, R.-T. | | Deposit date: | 2012-11-27 | | Release date: | 2013-04-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Structural and functional analyses of catalytic domain of GH10 xylanase from Thermoanaerobacterium saccharolyticum JW/SL-YS485
Proteins, 81, 2013
|
|
3W27
 
 | | The high-resolution crystal structure of TsXylA, intracellular xylanase from /Thermoanaerobacterium saccharolyticum JW/SL-YS485/: the complex of the E251A mutant with xylobiose | | Descriptor: | Glycoside hydrolase family 10, beta-D-xylopyranose-(1-4)-alpha-D-xylopyranose | | Authors: | Han, X, Gao, J, Shang, N, Huang, C.-H, Ko, T.-P, Zhu, Z, Wiegel, J, Shao, W, Guo, R.-T. | | Deposit date: | 2012-11-27 | | Release date: | 2013-04-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.41 Å) | | Cite: | Structural and functional analyses of catalytic domain of GH10 xylanase from Thermoanaerobacterium saccharolyticum JW/SL-YS485
Proteins, 81, 2013
|
|
8C0B
 | | CryoEM structure of Aspergillus nidulans UTP-glucose-1-phosphate uridylyltransferase | | Descriptor: | UTP--glucose-1-phosphate uridylyltransferase | | Authors: | Han, X, D Angelo, C, Otamendi, A, Cifuente, J.O, de Astigarraga, E, Ochoa-Lizarralde, B, Grininger, M, Routier, F.H, Guerin, M.E, Fuehring, J, Etxebeste, O, Connell, S.R. | | Deposit date: | 2022-12-16 | | Release date: | 2023-06-28 | | Last modified: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (3.98 Å) | | Cite: | CryoEM analysis of the essential native UDP-glucose pyrophosphorylase from Aspergillus nidulans reveals key conformations for activity regulation and function.
Mbio, 14, 2023
|
|
9IYI
 
 | | VLP structure of Chikungunya virus complexed with C34 Fab, 2f block. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, C34 Fab, ... | | Authors: | Han, X, Ji, C, Wang, F, Tian, S, Gao, F.G, Yan, J. | | Deposit date: | 2024-07-30 | | Release date: | 2025-11-12 | | Method: | ELECTRON MICROSCOPY (3.73 Å) | | Cite: | Neutralizing antibodies against Chikungunya virus and structural elucidation of their mechanism of action.
Nat Commun, 16, 2025
|
|
4QQR
 
 | |
9J6D
 
 | | Structure of Chikungunya virus infectious particles, 2f block. | | Descriptor: | CHIKV capsid protein, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Han, X, Ji, C, Wang, F, Tian, S, Gao, F.G, Yan, J. | | Deposit date: | 2024-08-15 | | Release date: | 2025-12-03 | | Method: | ELECTRON MICROSCOPY (3.11 Å) | | Cite: | Neutralizing antibodies against Chikungunya virus and structural elucidation of their mechanism of action.
Nat Commun, 16, 2025
|
|
9IXA
 
 | | Cryo-EM structure of chikungunya virus glycoprotein E1-E2 with C34 Fab. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, C34 Fab, Heavy Chain, ... | | Authors: | Han, X, Ji, C, Wang, F, Tian, S, Gao, F.G, Yan, J. | | Deposit date: | 2024-07-26 | | Release date: | 2025-11-19 | | Method: | ELECTRON MICROSCOPY (3.11 Å) | | Cite: | Neutralizing antibodies against Chikungunya virus and structural elucidation of their mechanism of action.
Nat Commun, 16, 2025
|
|
9IXI
 
 | | VLP structure of Chikungunya virus, 2f block. | | Descriptor: | CHIKV E1, CHIKV E2, CHIKV capsid protein | | Authors: | Han, X, Ji, C, Wang, F, Tian, S, Gao, F.G, Yan, J. | | Deposit date: | 2024-07-28 | | Release date: | 2025-11-19 | | Method: | ELECTRON MICROSCOPY (3.01 Å) | | Cite: | Neutralizing antibodies against Chikungunya virus and structural elucidation of their mechanism of action.
Nat Commun, 16, 2025
|
|
7F8V
 
 | | Cryo-EM structure of the cholecystokinin receptor CCKBR in complex with gastrin-17 and Gi | | Descriptor: | Gastrin-17, Gastrin/cholecystokinin type B receptor, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Zhang, X, He, C, Wang, M, Zhou, Q, Yang, D, Zhu, Y, Wu, B, Zhao, Q. | | Deposit date: | 2021-07-02 | | Release date: | 2021-10-13 | | Last modified: | 2022-02-16 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structures of the human cholecystokinin receptors bound to agonists and antagonists.
Nat.Chem.Biol., 17, 2021
|
|
