1DJH
 
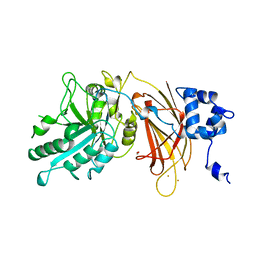 | | PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH BARIUM | | Descriptor: | ACETATE ION, BARIUM ION, PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C, ... | | Authors: | Essen, L.-O, Perisic, O, Williams, R.L. | | Deposit date: | 1996-09-25 | | Release date: | 1997-07-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A ternary metal binding site in the C2 domain of phosphoinositide-specific phospholipase C-delta1.
Biochemistry, 36, 1997
|
|
1DJZ
 
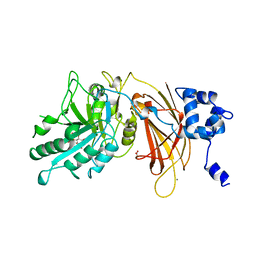 | | PHOSPHOINOSITIDE-SPECIFIC PHOSPHOLIPASE C-DELTA1 FROM RAT COMPLEXED WITH INOSITOL-4,5-BISPHOSPHATE | | Descriptor: | ACETATE ION, CALCIUM ION, D-MYO-INOSITOL-4,5-BISPHOSPHATE, ... | | Authors: | Essen, L.-O, Perisic, O, Williams, R.L. | | Deposit date: | 1996-08-24 | | Release date: | 1997-07-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural mapping of the catalytic mechanism for a mammalian phosphoinositide-specific phospholipase C.
Biochemistry, 36, 1997
|
|
5W5D
 
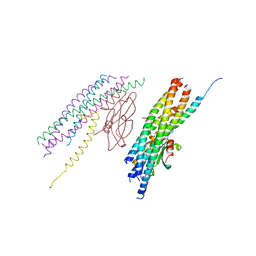 | |
5W5C
 
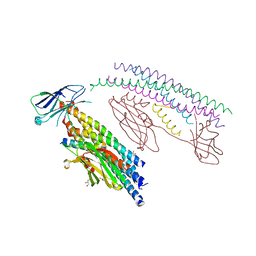 | |
4P42
 
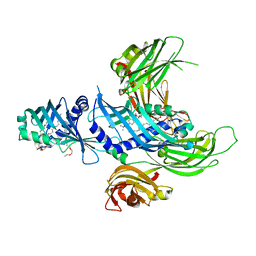 | | Extended-Synaptotagmin 2, SMP - C2A - C2B Domains | | Descriptor: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, 2-(2-{2-[2-(2-{2-[2-(2-{2-[4-(1,1,3,3-TETRAMETHYL-BUTYL)-PHENOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOX Y}-ETHOXY)-ETHANOL, Extended synaptotagmin-2 | | Authors: | Schauder, C.M, Wu, X, Reinisch, K.M. | | Deposit date: | 2014-03-10 | | Release date: | 2014-05-14 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Structure of a lipid-bound extended synaptotagmin indicates a role in lipid transfer.
Nature, 510, 2014
|
|
4QJ4
 
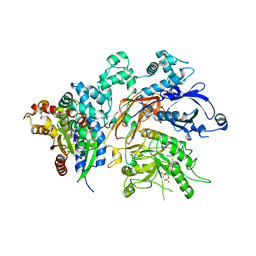 | | Structure of a fragment of human phospholipase C-beta3 delta472-569, bound to IP3 and in complex with Galphaq | | Descriptor: | 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase beta-3, CALCIUM ION, D-MYO-INOSITOL-1,4,5-TRIPHOSPHATE, ... | | Authors: | Lyon, A.M, Tesmer, J.J.G. | | Deposit date: | 2014-06-03 | | Release date: | 2014-10-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Molecular mechanisms of phospholipase C beta 3 autoinhibition.
Structure, 22, 2014
|
|
2CJS
 
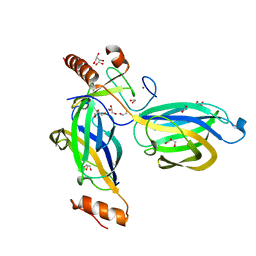 | | Structural Basis for a Munc13-1 Homodimer - Munc13-1 - RIM Heterodimer Switch: C2-domains as Versatile Protein-Protein Interaction Modules | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, REGULATING SYNAPTIC MEMBRANE EXOCYTOSIS PROTEIN 2, ... | | Authors: | Lu, J, Machius, M, Dulubova, I, Dai, H, Sudhof, T.C, Tomchick, D.R, Rizo, J. | | Deposit date: | 2006-04-06 | | Release date: | 2006-06-07 | | Last modified: | 2019-01-30 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Structural Basis for a Munc13-1 Homodimer to Munc13-1/Rim Heterodimer Switch.
Plos Biol., 4, 2006
|
|
7Z3J
 
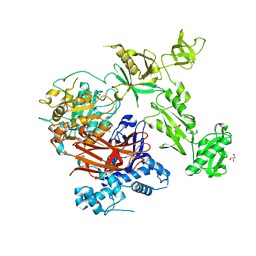 | | Structure of crystallisable rat Phospholipase C gamma 1 in complex with inositol 1,4,5-trisphosphate | | Descriptor: | 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase gamma-1, CALCIUM ION, D-MYO-INOSITOL-1,4,5-TRIPHOSPHATE, ... | | Authors: | Pinotsis, N, Bunney, T.D, Katan, M. | | Deposit date: | 2022-03-02 | | Release date: | 2022-07-20 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Characterization of the membrane interactions of phospholipase C gamma reveals key features of the active enzyme.
Sci Adv, 8, 2022
|
|
6UWA
 
 | | Mouse PKC C1B and C2 domains | | Descriptor: | CITRIC ACID, LEAD (II) ION, Protein kinase C, ... | | Authors: | Aggarwal, A, Mire, J, Sacchettini, J.C, Igumenova, T. | | Deposit date: | 2019-11-04 | | Release date: | 2020-12-09 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Mouse PKC C1B and C2 domains
To Be Published
|
|
7BL1
 
 | | human complex II-BATS bound to membrane-attached Rab5a-GTP | | Descriptor: | Beclin-1, GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Tremel, S, Morado, D.R, Kovtun, O, Williams, R.L, Briggs, J.A.G, Munro, S, Ohashi, Y, Bertram, J, Perisic, O. | | Deposit date: | 2021-01-17 | | Release date: | 2021-03-03 | | Last modified: | 2021-03-24 | | Method: | ELECTRON MICROSCOPY (9.8 Å) | | Cite: | Structural basis for VPS34 kinase activation by Rab1 and Rab5 on membranes.
Nat Commun, 12, 2021
|
|
6PBC
 
 | | Structural basis for the activation of PLC-gamma isozymes by phosphorylation and cancer-associated mutations | | Descriptor: | 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase gamma,1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase gamma-1, CALCIUM ION, SODIUM ION | | Authors: | Hajicek, N, Sondek, J. | | Deposit date: | 2019-06-13 | | Release date: | 2020-01-08 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Structural basis for the activation of PLC-gamma isozymes by phosphorylation and cancer-associated mutations.
Elife, 8, 2019
|
|
3M7F
 
 | | Crystal structure of the Nedd4 C2/Grb10 SH2 complex | | Descriptor: | E3 ubiquitin-protein ligase NEDD4, Growth factor receptor-bound protein 10 | | Authors: | Huang, Q, Szebenyi, M. | | Deposit date: | 2010-03-16 | | Release date: | 2010-10-27 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Basis for the Interaction between the Growth Factor-binding Protein GRB10 and the E3 Ubiquitin Ligase NEDD4.
J.Biol.Chem., 285, 2010
|
|
3PFQ
 
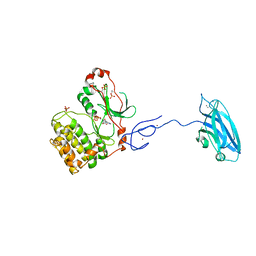 | | Crystal Structure and Allosteric Activation of Protein Kinase C beta II | | Descriptor: | CALCIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, Protein kinase C beta type, ... | | Authors: | Leonard, T.A, Rozycki, B, Saidi, L.F, Hummer, G, Hurley, J.H. | | Deposit date: | 2010-10-28 | | Release date: | 2011-02-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Crystal Structure and Allosteric Activation of Protein Kinase C beta II
Cell(Cambridge,Mass.), 144, 2011
|
|
7T7X
 
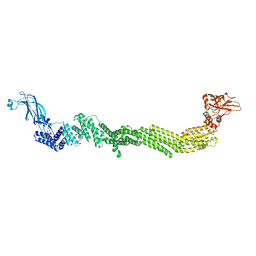 | |
7T7C
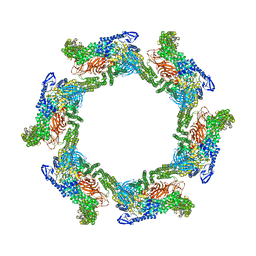 
 | |
7T7V
 
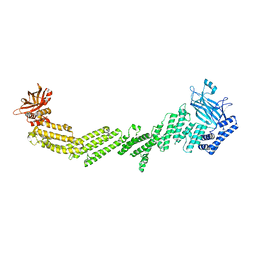 | |
7T81
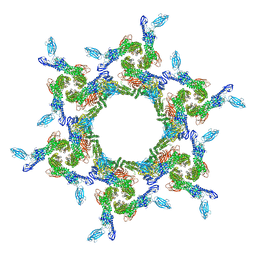 
 | |
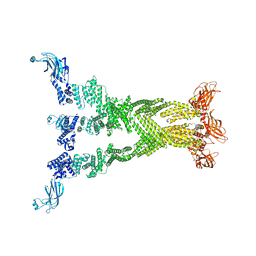
7T7R
 
 | |
7T8T
 
 | | CryoEM structure of PLCg1 | | Descriptor: | 1-phosphatidylinositol 4,5-bisphosphate phosphodiesterase gamma, CALCIUM ION | | Authors: | Endo-Streeter, S, Sondek, J. | | Deposit date: | 2021-12-17 | | Release date: | 2022-12-21 | | Method: | ELECTRON MICROSCOPY (3.68 Å) | | Cite: | CryoEM structure of PLCg1
To Be Published
|
|
2FJU
 
 | | Activated Rac1 bound to its effector phospholipase C beta 2 | | Descriptor: | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 2, 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, CALCIUM ION, ... | | Authors: | Jezyk, M.R, Snyder, J.T, Harden, T.K, Sondek, J. | | Deposit date: | 2006-01-03 | | Release date: | 2006-11-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of Rac1 bound to its effector phospholipase C-beta2.
Nat.Struct.Mol.Biol., 13, 2006
|
|
2K4A
 
 | |
2KI6
 
 | | The FGF1-S100A13-C2A hetero-hexameric complex structure: A component in the non-classical pathway for FGF1 secretion | | Descriptor: | Heparin-binding growth factor 1, Protein S100-A13, Synaptotagmin-1 | | Authors: | Krishna, S.M, Rani, S.G, Yu, C. | | Deposit date: | 2009-04-28 | | Release date: | 2010-03-09 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | The heterohexameric complex structure, a component in the non-classical pathway for fibroblast growth factor 1 (FGF1) secretion.
J.Biol.Chem., 285, 2010
|
|
5UE8
 
 | | The crystal structure of Munc13-1 C1C2BMUN domain | | Descriptor: | CHLORIDE ION, Protein unc-13 homolog A, ZINC ION | | Authors: | Tomchick, D.R, Rizo, J, Xu, J. | | Deposit date: | 2016-12-29 | | Release date: | 2017-02-15 | | Last modified: | 2018-03-28 | | Method: | X-RAY DIFFRACTION (3.35 Å) | | Cite: | Mechanistic insights into neurotransmitter release and presynaptic plasticity from the crystal structure of Munc13-1 C1C2BMUN.
Elife, 6, 2017
|
|
