3KRJ
 
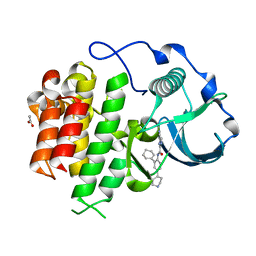 | | cFMS tyrosine kinase in complex with 4-Cyano-1H-imidazole-2-carboxylic acid (2-cyclohex-1-enyl-4-piperidin-4-yl-phenyl)-amide | | Descriptor: | 4-cyano-N-(2-cyclohex-1-en-1-yl-4-piperidin-4-ylphenyl)-1H-imidazole-2-carboxamide, ACETATE ION, Macrophage colony-stimulating factor 1 receptor, ... | | Authors: | Schubert, C. | | Deposit date: | 2009-11-18 | | Release date: | 2010-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Optimization of a Potent Class of Arylamide Colony-Stimulating Factor-1 Receptor Inhibitors Leading to Anti-inflammatory Clinical Candidate 4-Cyano-N-[2-(1-cyclohexen-1-yl)-4-[1-[(dimethylamino)acetyl]-4-piperidinyl]phenyl]-1H-imidazole-2-carboxamide (JNJ-28312141).
J.Med.Chem., 54, 2011
|
|
3KRL
 
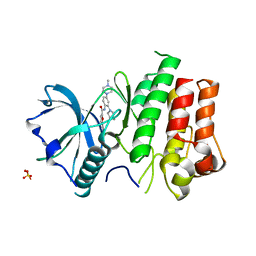 | | cFMS Tyrosine kinase in complex with 5-Cyano-furan-2-carboxylic acid [4-(4-methyl-piperazin-1-yl)-2-piperidin-1-yl-phenyl]-amide | | Descriptor: | 5-cyano-N-[4-(4-methylpiperazin-1-yl)-2-piperidin-1-ylphenyl]furan-2-carboxamide, Macrophage colony-stimulating factor 1 receptor, Basic fibroblast growth factor receptor 1, ... | | Authors: | Schubert, C. | | Deposit date: | 2009-11-18 | | Release date: | 2010-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Optimization of a Potent Class of Arylamide Colony-Stimulating Factor-1 Receptor Inhibitors Leading to Anti-inflammatory Clinical Candidate 4-Cyano-N-[2-(1-cyclohexen-1-yl)-4-[1-[(dimethylamino)acetyl]-4-piperidinyl]phenyl]-1H-imidazole-2-carboxamide (JNJ-28312141).
J.Med.Chem., 54, 2011
|
|
3PE6
 
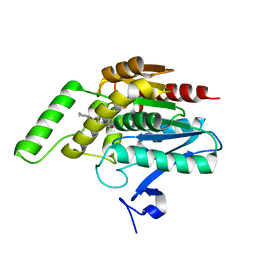 | |
1G4M
 
 | | CRYSTAL STRUCTURE OF BOVINE BETA-ARRESTIN 1 | | Descriptor: | BETA-ARRESTIN1 | | Authors: | Schubert, C, Han, M. | | Deposit date: | 2000-10-27 | | Release date: | 2001-10-03 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of beta-arrestin at 1.9 A: possible mechanism of receptor binding and membrane Translocation.
Structure, 9, 2001
|
|
1G4R
 
 | | CRYSTAL STRUCTURE OF BOVINE BETA-ARRESTIN 1 | | Descriptor: | BETA-ARRESTIN 1 | | Authors: | Schubert, C, Han, M. | | Deposit date: | 2000-10-27 | | Release date: | 2001-10-03 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of beta-arrestin at 1.9 A: possible mechanism of receptor binding and membrane Translocation.
Structure, 9, 2001
|
|
5C37
 
 | | Structure of the beta-ketoacyl reductase domain of human fatty acid synthase bound to a spiro-imidazolone inhibitor | | Descriptor: | 6-{[(3R)-1-(cyclopropylcarbonyl)pyrrolidin-3-yl]methyl}-5-[4-(1-methyl-1H-indazol-5-yl)phenyl]-4,6-diazaspiro[2.4]hept-4-en-7-one, CHLORIDE ION, Fatty acid synthase, ... | | Authors: | Schubert, C, Milligan, C.M, Vo, K, Grasberger, B. | | Deposit date: | 2015-06-17 | | Release date: | 2016-06-22 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Design and Synthesis of a Series of Bioavailable Fatty Acid Synthase (FASN) KR Domain Inhibitors for Cancer Therapy
to be published
|
|
3BEA
 
 | | cFMS tyrosine kinase (tie2 KID) in complex with a pyrimidinopyridone inhibitor | | Descriptor: | 8-(2,3-dihydro-1H-inden-5-yl)-2-({4-[(3R,5S)-3,5-dimethylpiperazin-1-yl]phenyl}amino)-5-oxo-5,8-dihydropyrido[2,3-d]pyrimidine-6-carboxamide, Macrophage colony-stimulating factor 1 receptor, SULFATE ION | | Authors: | Schubert, C. | | Deposit date: | 2007-11-16 | | Release date: | 2008-07-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Design and synthesis of a pyrido[2,3-d]pyrimidin-5-one class of anti-inflammatory FMS inhibitors.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
3DPK
 
 | | cFMS tyrosine kinase in complex with a pyridopyrimidinone inhibitor | | Descriptor: | 8-cyclohexyl-N-methoxy-5-oxo-2-{[4-(2-pyrrolidin-1-ylethyl)phenyl]amino}-5,8-dihydropyrido[2,3-d]pyrimidine-6-carboxamide, Macrophage colony-stimulating factor 1 receptor, Fibroblast growth factor receptor 1, ... | | Authors: | Schubert, C. | | Deposit date: | 2008-07-08 | | Release date: | 2009-02-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Pyrido[2,3-d]pyrimidin-5-ones: a novel class of antiinflammatory macrophage colony-stimulating factor-1 receptor inhibitors
J.Med.Chem., 52, 2009
|
|
2I0V
 
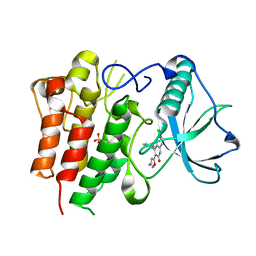 | | c-FMS tyrosine kinase in complex with a quinolone inhibitor | | Descriptor: | 6-CHLORO-3-(3-METHYLISOXAZOL-5-YL)-4-PHENYLQUINOLIN-2(1H)-ONE, SULFATE ION, cFMS tyrosine kinase | | Authors: | Schubert, C, Schalk-Hihi, C. | | Deposit date: | 2006-08-11 | | Release date: | 2006-11-28 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the tyrosine kinase domain of colony-stimulating factor-1 receptor (cFMS) in complex with two inhibitors.
J.Biol.Chem., 282, 2007
|
|
2I1M
 
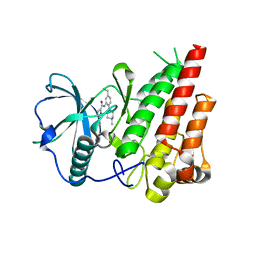 | | cFMS tyrosine kinase (tie2 KID) in complex with an arylamide inhibitor | | Descriptor: | 5-CYANO-FURAN-2-CARBOXYLIC ACID [5-HYDROXYMETHYL-2-(4-METHYL-PIPERIDIN-1-YL)-PHENYL]-AMIDE, Macrophage colony-stimulating factor 1 receptor | | Authors: | Schubert, C, Schalk-Hihi, C. | | Deposit date: | 2006-08-14 | | Release date: | 2006-11-28 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of the tyrosine kinase domain of colony-stimulating factor-1 receptor (cFMS) in complex with two inhibitors.
J.Biol.Chem., 282, 2007
|
|
2GV2
 
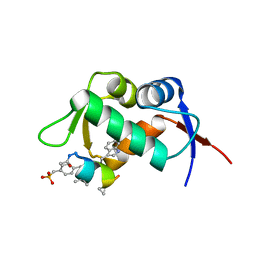 | | MDM2 in complex with an 8-mer p53 peptide analogue | | Descriptor: | 8-MER P53 PEPTIDE ANALOGUE, E3 ubiquitin-protein ligase Mdm2 | | Authors: | Schubert, C, Sakurai, K. | | Deposit date: | 2006-05-02 | | Release date: | 2006-09-05 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystallographic Analysis of an 8-mer p53 Peptide Analogue Complexed with MDM2.
J.Am.Chem.Soc., 128, 2006
|
|
2I0Y
 
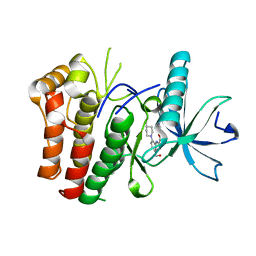 | | cFMS tyrosine kinase (FGF KID) in complex with an arylamide inhibitor | | Descriptor: | 5-CYANO-FURAN-2-CARBOXYLIC ACID [5-HYDROXYMETHYL-2-(4-METHYL-PIPERIDIN-1-YL)-PHENYL]-AMIDE, cFMS Tyrosine Kinase | | Authors: | Schubert, C, Schalk-Hihi, C. | | Deposit date: | 2006-08-11 | | Release date: | 2006-11-28 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the tyrosine kinase domain of colony-stimulating factor-1 receptor (cFMS) in complex with two inhibitors.
J.Biol.Chem., 282, 2007
|
|
4ATX
 
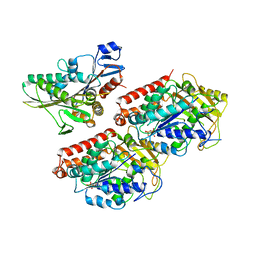 | | Rigor kinesin motor domain with an ordered neck-linker, docked on tubulin dimer, modelled into the 8A cryo-EM map of doublecortin- microtubules decorated with kinesin | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, KINESIN-1 HEAVY CHAIN, ... | | Authors: | Liu, J.S, Schubert, C.R, Fu, X, Fourniol, F.J, Jaiswal, J.K, Houdusse, A, Stultz, C.M, Moores, C.A, Walsh, C.A. | | Deposit date: | 2012-05-10 | | Release date: | 2012-09-26 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (8.2 Å) | | Cite: | Molecular Basis for Specific Regulation of Neuronal Kinesin- 3 Motors by Doublecortin Family Proteins.
Mol.Cell, 47, 2012
|
|
1CF1
 
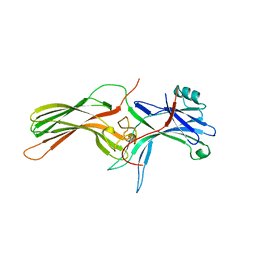 | | ARRESTIN FROM BOVINE ROD OUTER SEGMENTS | | Descriptor: | PROTEIN (ARRESTIN) | | Authors: | Hirsch, J.A, Schubert, C, Gurevich, V.V, Sigler, P.B. | | Deposit date: | 1999-03-23 | | Release date: | 1999-04-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The 2.8 A crystal structure of visual arrestin: a model for arrestin's regulation.
Cell(Cambridge,Mass.), 97, 1999
|
|
4ATU
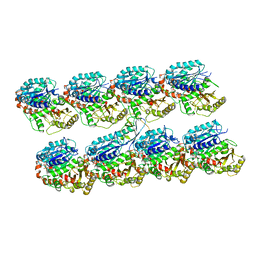 
 | | Human doublecortin N-DC repeat plus linker, and tubulin (2XRP) docked into an 8A cryo-EM map of doublecortin-stabilised microtubules reconstructed in absence of kinesin | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, NEURONAL MIGRATION PROTEIN DOUBLECORTIN, ... | | Authors: | Liu, J.S, Schubert, C.R, Fu, X, Fourniol, F.J, Jaiswal, J.K, Houdusse, A, Stultz, C.M, Moores, C.A, Walsh, C.A. | | Deposit date: | 2012-05-09 | | Release date: | 2012-09-26 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (8.3 Å) | | Cite: | Molecular Basis for Specific Regulation of Neuronal Kinesin- 3 Motors by Doublecortin Family Proteins.
Mol.Cell, 47, 2012
|
|
1T4E
 
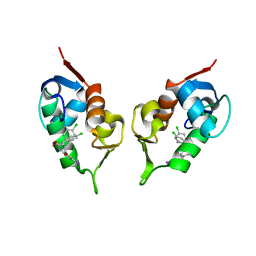 | | Structure of Human MDM2 in complex with a Benzodiazepine Inhibitor | | Descriptor: | (4-CHLOROPHENYL)[3-(4-CHLOROPHENYL)-7-IODO-2,5-DIOXO-1,2,3,5-TETRAHYDRO-4H-1,4-BENZODIAZEPIN-4-YL]ACETIC ACID, Ubiquitin-protein ligase E3 Mdm2 | | Authors: | Grasberger, B.L, Schubert, C, Koblish, H.K, Carver, T.E, Franks, C.F, Zhao, S.Y, Lu, T, LaFrance, L.V, Parks, D.J. | | Deposit date: | 2004-04-29 | | Release date: | 2005-02-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Discovery and cocrystal structure of benzodiazepinedione HDM2 antagonists that activate p53 in cells
J.Med.Chem., 48, 2005
|
|
1T4F
 
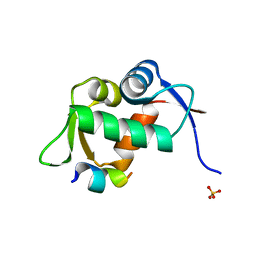 | | Structure of human MDM2 in complex with an optimized p53 peptide | | Descriptor: | SULFATE ION, Ubiquitin-protein ligase E3 Mdm2, optimized p53 peptide | | Authors: | Grasberger, B.L, Schubert, C, Koblish, H.K, Carver, T.E, Franks, C.F, Zhao, S.Y, Lu, T, LaFrance, L.V, Parks, D.J. | | Deposit date: | 2004-04-29 | | Release date: | 2005-02-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Discovery and cocrystal structure of benzodiazepinedione HDM2 antagonists that activate p53 in cells
J.Med.Chem., 48, 2005
|
|
5K9R
 
 | | PDE10a with imidazopyrazine inhibitor | | Descriptor: | 4-[5-[1-(2-methoxyethyl)pyrazol-4-yl]-2-(quinolin-2-yloxymethyl)imidazo[1,2-a]pyrazin-8-yl]morpholine, MAGNESIUM ION, ZINC ION, ... | | Authors: | Gibbs, A.G, Schubert, C. | | Deposit date: | 2016-06-01 | | Release date: | 2016-08-24 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Discovery of novel potent imidazo[1,2-b]pyridazine PDE10a inhibitors.
Bioorg.Med.Chem.Lett., 26, 2016
|
|
