6CFN
 
 | |
3LUH
 
 | |
3LUG
 
 | |
3LUC
 
 | |
3LUJ
 
 | |
3QX9
 
 | | Crystal structure of MID domain from hAGO2 in complex with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Protein argonaute-2 | | Authors: | Frank, F, Fabian, M.R, Stepinski, J, Jemielity, J, Darzynkiewicz, E, Sonenberg, N, Nagar, B. | | Deposit date: | 2011-03-01 | | Release date: | 2011-04-20 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural analysis of 5'-mRNA-cap interactions with the human AGO2 MID domain.
Embo Rep., 12, 2011
|
|
3QX8
 
 | | Crystal structure of MID domain from hAGO2 in complex with m7GpppG | | Descriptor: | 7-METHYL-GUANOSINE-5'-TRIPHOSPHATE-5'-GUANOSINE, Protein argonaute-2 | | Authors: | Frank, F, Fabian, M.R, Stepinski, J, Jemielity, J, Darzynkiewicz, E, Sonenberg, N, Nagar, B. | | Deposit date: | 2011-03-01 | | Release date: | 2011-04-20 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural analysis of 5'-mRNA-cap interactions with the human AGO2 MID domain.
Embo Rep., 12, 2011
|
|
4G0X
 
 | | Crystal Structure of Arabidopsis thaliana AGO1 MID domain | | Descriptor: | Protein argonaute 1, SULFATE ION | | Authors: | Frank, F, Hauver, J, Sonenberg, N, Nagar, B. | | Deposit date: | 2012-07-10 | | Release date: | 2012-07-25 | | Last modified: | 2012-09-12 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Arabidopsis Argonaute MID domains use their nucleotide specificity loop to sort small RNAs.
Embo J., 31, 2012
|
|
4G0P
 
 | | Crystal Structure of Arabidopsis thaliana AGO1 MID domain in complex with UMP | | Descriptor: | Protein argonaute 1, URIDINE-5'-MONOPHOSPHATE | | Authors: | Frank, F, Hauver, J, Sonenberg, N, Nagar, B. | | Deposit date: | 2012-07-09 | | Release date: | 2012-07-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Arabidopsis Argonaute MID domains use their nucleotide specificity loop to sort small RNAs.
Embo J., 31, 2012
|
|
4G0Y
 
 | | Crystal structure of Arabidopsis AGO1 in complex with AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, Protein argonaute 1 | | Authors: | Frank, F, Hauver, J, Sonenberg, N, Nagar, B. | | Deposit date: | 2012-07-10 | | Release date: | 2012-07-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Arabidopsis Argonaute MID domains use their nucleotide specificity loop to sort small RNAs.
Embo J., 31, 2012
|
|
4G0O
 
 | | Crystal structure of Arabidopsis thaliana AGO5 MID domain | | Descriptor: | Protein argonaute 5, SULFATE ION | | Authors: | Frank, F, Hauver, J, Sonenberg, N, Nagar, B. | | Deposit date: | 2012-07-09 | | Release date: | 2012-07-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.186 Å) | | Cite: | Arabidopsis Argonaute MID domains use their nucleotide specificity loop to sort small RNAs.
Embo J., 31, 2012
|
|
4G0Z
 
 | | Crystal structure of Arabidopsis thaliana AGO1 MID domain in complex with GMP | | Descriptor: | GUANOSINE-5'-MONOPHOSPHATE, Protein argonaute 1 | | Authors: | Frank, F, Hauver, J, Sonenberg, N, Nagar, B. | | Deposit date: | 2012-07-10 | | Release date: | 2012-07-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Arabidopsis Argonaute MID domains use their nucleotide specificity loop to sort small RNAs.
Embo J., 31, 2012
|
|
4G0M
 
 | | Crystal structure of Arabidopsis thaliana AGO2 MID domain | | Descriptor: | Protein argonaute 2, SULFATE ION | | Authors: | Frank, F, Hauver, J, Sonenberg, N, Nagar, B. | | Deposit date: | 2012-07-09 | | Release date: | 2012-07-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.306 Å) | | Cite: | Arabidopsis Argonaute MID domains use their nucleotide specificity loop to sort small RNAs.
Embo J., 31, 2012
|
|
4G0Q
 
 | | Crystal structure of Arabidopsis thaliana AGO1 MID domain in complex with CMP | | Descriptor: | CYTIDINE-5'-MONOPHOSPHATE, Protein argonaute 1 | | Authors: | Frank, F, Hauver, J, Sonenberg, N, Nagar, B. | | Deposit date: | 2012-07-09 | | Release date: | 2012-07-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Arabidopsis Argonaute MID domains use their nucleotide specificity loop to sort small RNAs.
Embo J., 31, 2012
|
|
3LUD
 
 | |
3LUK
 
 | |
4IUL
 
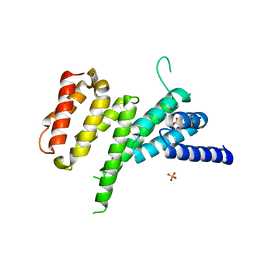 | | MIF4G domain of DAP5 | | Descriptor: | Eukaryotic translation initiation factor 4 gamma 2, SULFATE ION | | Authors: | Frank, F, Virgili, G, Feoktistova, K, Sawicki, M, Sonenberg, N, Fraser, C, Nagar, B. | | Deposit date: | 2013-01-21 | | Release date: | 2013-03-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural Analysis of the DAP5 MIF4G Domain and Its Interaction with eIF4A.
Structure, 21, 2013
|
|
4J8S
 
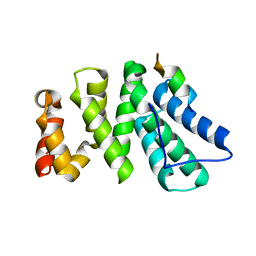 | | Crystal structure of human CNOT1 MIF4G domain in complex with a TTP peptide | | Descriptor: | CCR4-NOT transcription complex subunit 1, Tristetraprolin | | Authors: | Frank, F, Fabian, M.R, Rouya, C, Siddiqui, N, Lai, W.S, Karetnikov, A, Blackshear, P.J, Sonenberg, N, Nagar, B. | | Deposit date: | 2013-02-14 | | Release date: | 2013-05-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural basis for the recruitment of the human CCR4-NOT deadenylase complex by tristetraprolin.
Nat.Struct.Mol.Biol., 20, 2013
|
|
2OIQ
 
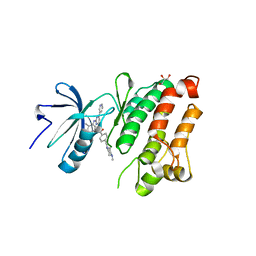 | | Crystal Structure of chicken c-Src kinase domain in complex with the cancer drug imatinib. | | Descriptor: | 4-(4-METHYL-PIPERAZIN-1-YLMETHYL)-N-[4-METHYL-3-(4-PYRIDIN-3-YL-PYRIMIDIN-2-YLAMINO)-PHENYL]-BENZAMIDE, GLYCEROL, Proto-oncogene tyrosine-protein kinase Src | | Authors: | Seeliger, M.A, Nagar, B, Frank, F, Cao, X, Henderson, M.N, Kuriyan, J. | | Deposit date: | 2007-01-11 | | Release date: | 2007-03-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | c-Src Binds to the Cancer Drug Imatinib with an Inactive Abl/c-Kit Conformation and a Distributed Thermodynamic Penalty.
Structure, 15, 2007
|
|
7A0Y
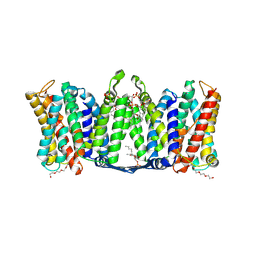 
 | | Structure of dimeric sodium proton antiporter NhaA K300R variant, at pH 8.2, crystallized with chimeric Fab antibodies | | Descriptor: | 1,2-DIACYL-GLYCEROL-3-SN-PHOSPHATE, 1,2-Distearoyl-sn-glycerophosphoethanolamine, CARDIOLIPIN, ... | | Authors: | Fippel, A, Mir, S.H, Lentes, C.J, Wirth, C, Hunte, C. | | Deposit date: | 2020-08-11 | | Release date: | 2022-02-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Molecular determinants for substrate uptake in electrogenic sodium/proton antiporters
To Be Published
|
|
7A0X
 
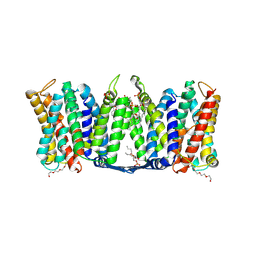 | | Structure of dimeric sodium proton antiporter NhaA, at pH 6.0, crystallized with chimeric Fab antibodies | | Descriptor: | 1,2-DIACYL-GLYCEROL-3-SN-PHOSPHATE, 1,2-Distearoyl-sn-glycerophosphoethanolamine, CALCIUM ION, ... | | Authors: | Fippel, A, Lentes, C.J, Mir, S.H, Wirth, C, Hunte, C. | | Deposit date: | 2020-08-11 | | Release date: | 2022-02-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Molecular determinants for substrate uptake in electrogenic sodium/proton antiporters
To Be Published
|
|
7A0W
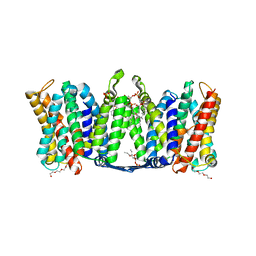 
 | | Structure of dimeric sodium proton antiporter NhaA, at pH 8.5, crystallized with chimeric Fab antibodies | | Descriptor: | 1,2-DIACYL-GLYCEROL-3-SN-PHOSPHATE, CARDIOLIPIN, CHLORIDE ION, ... | | Authors: | Fippel, A, Gross, N.M, Mir, S.H, Wirth, C, Hunte, C. | | Deposit date: | 2020-08-11 | | Release date: | 2022-02-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Molecular determinants for substrate uptake in electrogenic sodium/proton antiporters
To Be Published
|
|
8GDR
 
 | |
6EU6
 
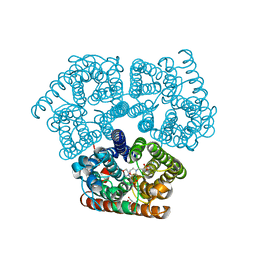 | | Sensor Amt Protein | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, AMMONIUM ION, Ammonium transporter, ... | | Authors: | Pflueger, T, Hernandez, C, Andrade, S.L.A. | | Deposit date: | 2017-10-28 | | Release date: | 2018-01-24 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Signaling ammonium across membranes through an ammonium sensor histidine kinase.
Nat Commun, 9, 2018
|
|
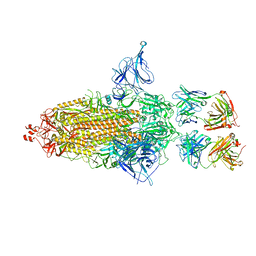
7U0P
 
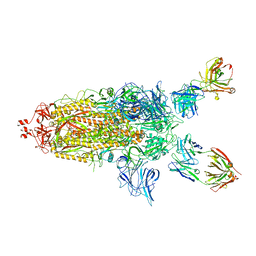 | | SARS-Cov2 S protein structure in complex with neutralizing monoclonal antibody 002-S21F2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Patel, A, Ortlund, E. | | Deposit date: | 2022-02-18 | | Release date: | 2022-08-10 | | Last modified: | 2023-02-22 | | Method: | ELECTRON MICROSCOPY (3.76 Å) | | Cite: | Structural insights for neutralization of Omicron variants BA.1, BA.2, BA.4, and BA.5 by a broadly neutralizing SARS-CoV-2 antibody.
Sci Adv, 8, 2022
|
|
