8YLE
 
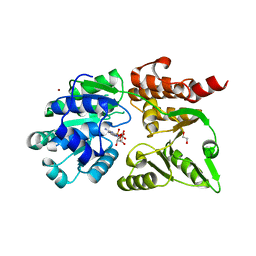 | | Crystal structure of Werner syndrome helicase complexed with AMP-PCP | | Descriptor: | 1,2-ETHANEDIOL, Bifunctional 3'-5' exonuclease/ATP-dependent helicase WRN, PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER, ... | | Authors: | Yang, Y, Fu, L, Sun, X, Cheng, H, Chen, R. | | Deposit date: | 2024-03-06 | | Release date: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Structure of werner syndrome helicase complexed with AMP-PCP at 1.86 Angstroms resolution.
To Be Published
|
|
5UFU
 
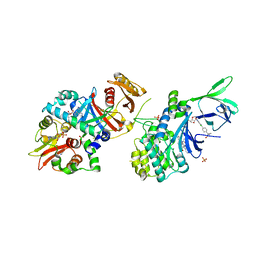 | | Structure of AMPK bound to activator | | Descriptor: | 1,4:3,6-dianhydro-2-O-(6-chloro-5-{4-[1-(hydroxymethyl)cyclopropyl]phenyl}-1H-benzimidazol-2-yl)-D-mannitol, 5'-AMP-activated protein kinase catalytic subunit alpha-1, 5'-AMP-activated protein kinase subunit beta-1, ... | | Authors: | Calabrese, M.F, Kurumbail, R.G. | | Deposit date: | 2017-01-05 | | Release date: | 2017-05-17 | | Method: | X-RAY DIFFRACTION (3.45 Å) | | Cite: | Activation of Skeletal Muscle AMPK Promotes Glucose Disposal and Glucose Lowering in Non-human Primates and Mice.
Cell Metab., 25, 2017
|
|
4UN0
 
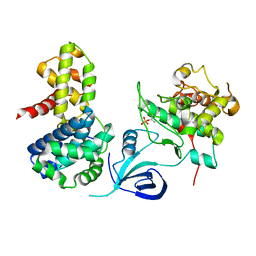 | | Crystal structure of the human CDK12-cyclinK complex | | Descriptor: | CYCLIN-DEPENDENT KINASE 12, CYCLIN-K | | Authors: | Dixon Clarke, S.E, Elkins, J.M, Pike, A.C.W, Chaikuad, A, Goubin, S, Krojer, T, Sorrell, F.J, Nowak, R, Williams, E, Kopec, J, Mahajan, R.P, Burgess-Brown, N, Carpenter, E.P, Knapp, S, von Delft, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Bullock, A. | | Deposit date: | 2014-05-22 | | Release date: | 2014-06-04 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Structures of the Cdk12/Cyck Complex with AMP-Pnp Reveal a Flexible C-Terminal Kinase Extension Important for ATP Binding.
Sci.Rep., 5, 2015
|
|
8C4J
 
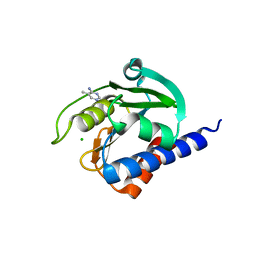 | | CdaA-compound 4 complex | | Descriptor: | 5-methylpyrimidin-4-amine, CHLORIDE ION, Diadenylate cyclase | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4P
 
 | | CdaA-compound 7 complex | | Descriptor: | CHLORIDE ION, Diadenylate cyclase, ~{N}-[5-(2-azanyl-1,3-thiazol-4-yl)-4-methyl-1,3-thiazol-2-yl]ethanamide | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4O
 
 | | CdaA-ATP complex | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CHLORIDE ION, Diadenylate cyclase, ... | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4M
 
 | | CdaA-Adenosine complex | | Descriptor: | ADENOSINE, CHLORIDE ION, Diadenylate cyclase | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4Q
 
 | | CdaA-Apo | | Descriptor: | CHLORIDE ION, Diadenylate cyclase | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4R
 
 | | CdaA-adenine complex | | Descriptor: | ADENINE, CHLORIDE ION, Diadenylate cyclase | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
7P9N
 
 | | Crystal structure of CD73 in complex with AMP in the open form | | Descriptor: | 5'-nucleotidase, ADENOSINE MONOPHOSPHATE, CALCIUM ION, ... | | Authors: | Scaletti, E.R, Strater, N. | | Deposit date: | 2021-07-27 | | Release date: | 2021-10-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Substrate binding modes of purine and pyrimidine nucleotides to human ecto-5'-nucleotidase (CD73) and inhibition by their bisphosphonic acid derivatives.
Purinergic Signal, 17, 2021
|
|
4YP1
 
 | | Misting with CDA | | Descriptor: | (2R,3R,3aS,5R,7aR,9R,10R,10aS,12R,14aR)-2,9-bis(6-amino-9H-purin-9-yl)octahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8 ]tetraoxadiphosphacyclododecine-3,5,10,12-tetrol 5,12-dioxide, Stimulator of interferon genes protein | | Authors: | Chin, K.H, Chen, C.K, Tu, Z.I, Chou, S.H. | | Deposit date: | 2015-03-12 | | Release date: | 2015-10-07 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural Insights into the Distinct Binding Mode of Cyclic Di-AMP with SaCpaA_RCK.
Biochemistry, 54, 2015
|
|
5VYW
 
 | | Crystal structure of Lactococcus lactis pyruvate carboxylase | | Descriptor: | BIOTIN, MANGANESE (II) ION, Pyruvate carboxylase | | Authors: | Choi, P.H, Tong, L. | | Deposit date: | 2017-05-26 | | Release date: | 2017-08-16 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural and functional studies of pyruvate carboxylase regulation by cyclic di-AMP in lactic acid bacteria.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
7Y0D
 
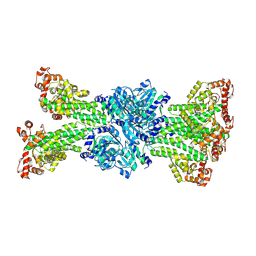 | |
6SHQ
 
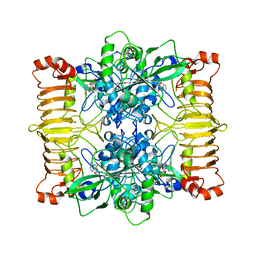 | | Escherichia coli AGPase in complex with AMP. Symmetry C2 | | Descriptor: | ADENOSINE MONOPHOSPHATE, Glucose-1-phosphate adenylyltransferase | | Authors: | Cifuente, J.O, Comino, N, D'Angelo, C, Marina, A, Gil-Carton, D, Albesa-Jove, D, Guerin, M.E. | | Deposit date: | 2019-08-07 | | Release date: | 2020-02-05 | | Last modified: | 2021-07-21 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The allosteric control mechanism of bacterial glycogen biosynthesis disclosed by cryoEM.
Curr Res Struct Biol, 2, 2020
|
|
6SI8
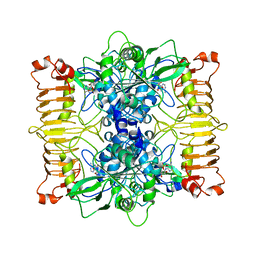 
 | | Escherichia coli AGPase in complex with AMP. | | Descriptor: | ADENOSINE MONOPHOSPHATE, Glucose-1-phosphate adenylyltransferase | | Authors: | Cifuente, J.O, Comino, N, D'Angelo, C, Marina, A, Gil-Carton, D, Albesa-Jove, D, Guerin, M.E. | | Deposit date: | 2019-08-09 | | Release date: | 2020-02-05 | | Last modified: | 2021-07-21 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | The allosteric control mechanism of bacterial glycogen biosynthesis disclosed by cryoEM.
Curr Res Struct Biol, 2, 2020
|
|
7OJR
 
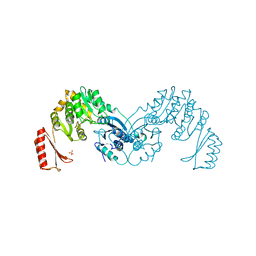 | |
7OML
 
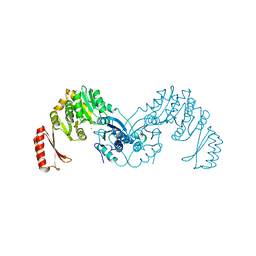 | |
8EPS
 
 | |
8G0V
 
 | |
8G0R
 
 | |
8G0U
 
 | |
8G0T
 
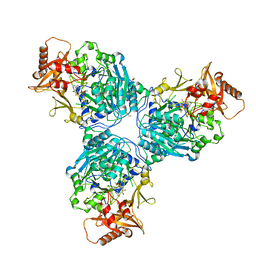 | |
8G0S
 
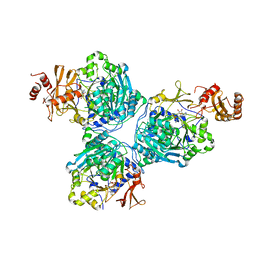 | |
4YB6
 
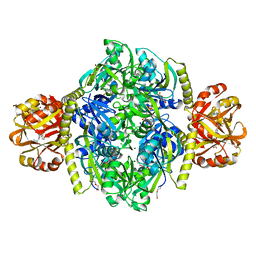 | | Adenosine triphosphate phosphoribosyltransferase from Campylobacter jejuni in complex with the inhibitors AMP and histidine | | Descriptor: | ADENOSINE MONOPHOSPHATE, ATP phosphoribosyltransferase, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Mittelstaedt, G, Moggre, G.-J, Parker, E.J. | | Deposit date: | 2015-02-18 | | Release date: | 2016-03-09 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Campylobacter jejuni adenosine triphosphate phosphoribosyltransferase is an active hexamer that is allosterically controlled by the twisting of a regulatory tail.
Protein Sci., 25, 2016
|
|
7XMV
 
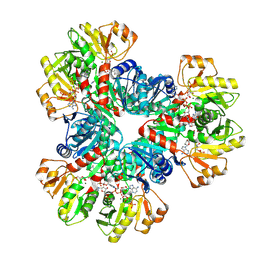 | | E.coli phosphoribosylpyrophosphate (PRPP) synthetase type A(AMP/ADP) filament bound with ADP, AMP and R5P | | Descriptor: | 5-O-phosphono-alpha-D-ribofuranose, ADENOSINE MONOPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Hu, H.H, Lu, G.M, Chang, C.C, Liu, J.L. | | Deposit date: | 2022-04-27 | | Release date: | 2022-06-29 | | Last modified: | 2022-07-06 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Filamentation modulates allosteric regulation of PRPS.
Elife, 11, 2022
|
|
