7PNT
 
 | | Assembly intermediate of mouse mitochondrial ribosome small subunit without mS37 in complex with RbfA and Tfb1m | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Itoh, Y, Khawaja, A, Laptev, I, Sergiev, P, Rorbach, J, Amunts, A. | | Deposit date: | 2021-09-08 | | Release date: | 2022-06-15 | | Last modified: | 2024-01-17 | | Method: | ELECTRON MICROSCOPY (3.19 Å) | | Cite: | Mechanism of mitoribosomal small subunit biogenesis and preinitiation.
Nature, 606, 2022
|
|
7PNW
 
 | | Mouse mitochondrial ribosome small subunit lacking m5U modification | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Itoh, Y, Khawaja, A, Laptev, I, Sergiev, P, Rorbach, J, Amunts, A. | | Deposit date: | 2021-09-08 | | Release date: | 2022-06-15 | | Last modified: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (3.09 Å) | | Cite: | Mechanism of mitoribosomal small subunit biogenesis and preinitiation.
Nature, 606, 2022
|
|
7PNU
 
 | | Assembly intermediate of mouse mitochondrial ribosome small subunit without mS37 in complex with RbfA inward conformation | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Itoh, Y, Khawaja, A, Laptev, I, Sergiev, P, Rorbach, J, Amunts, A. | | Deposit date: | 2021-09-08 | | Release date: | 2022-06-15 | | Last modified: | 2024-01-17 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Mechanism of mitoribosomal small subunit biogenesis and preinitiation.
Nature, 606, 2022
|
|
7PNV
 
 | | Assembly intermediate of mouse mitochondrial ribosome small subunit without mS37 in complex with RbfA and Mettl15 | | Descriptor: | 12S mitochondrial rRNA, 12S rRNA N4-methylcytidine methyltransferase, 28S ribosomal protein S10, ... | | Authors: | Itoh, Y, Khawaja, A, Laptev, I, Sergiev, P, Rorbach, J, Amunts, A. | | Deposit date: | 2021-09-08 | | Release date: | 2022-06-15 | | Last modified: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Mechanism of mitoribosomal small subunit biogenesis and preinitiation.
Nature, 606, 2022
|
|
3NWA
 
 | | Glycoprotein B from Herpes simplex virus type 1, W174R mutant, low-pH | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Envelope glycoprotein B, MESO-ERYTHRITOL | | Authors: | Stampfer, S.D, Lou, H, Cohen, G.H, Eisenberg, R.J, Heldwein, E.E. | | Deposit date: | 2010-07-09 | | Release date: | 2010-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2629 Å) | | Cite: | Structural basis of local, pH-dependent conformational changes in glycoprotein B from herpes simplex virus type 1.
J.Virol., 84, 2010
|
|
3NWF
 
 | | Glycoprotein B from Herpes simplex virus type 1, low-pH | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Envelope glycoprotein B, MESO-ERYTHRITOL, ... | | Authors: | Stampfer, S.D, Lou, H, Cohen, G.H, Eisenberg, R.J, Heldwein, E.E. | | Deposit date: | 2010-07-09 | | Release date: | 2010-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural basis of local, pH-dependent conformational changes in glycoprotein B from herpes simplex virus type 1.
J.Virol., 84, 2010
|
|
3NW8
 
 | | Glycoprotein B from Herpes simplex virus type 1, Y179S mutant, high-pH | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | Authors: | Stampfer, S.D, Lou, H, Cohen, G.H, Eisenberg, R.J, Heldwein, E.E. | | Deposit date: | 2010-07-09 | | Release date: | 2010-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.75915 Å) | | Cite: | Structural basis of local, pH-dependent conformational changes in glycoprotein B from herpes simplex virus type 1.
J.Virol., 84, 2010
|
|
3NWD
 
 | | Glycoprotein B from Herpes simplex virus type 1, Y179S mutant, low-pH | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | Authors: | Stampfer, S.D, Lou, H, Cohen, G.H, Eisenberg, R.J, Heldwein, E.E. | | Deposit date: | 2010-07-09 | | Release date: | 2010-12-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8803 Å) | | Cite: | Structural basis of local, pH-dependent conformational changes in glycoprotein B from herpes simplex virus type 1.
J.Virol., 84, 2010
|
|
7QEJ
 
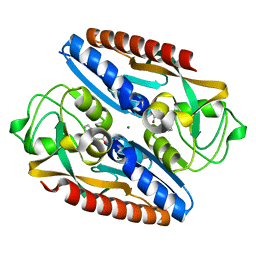 | | Structure of the ligand binding domain of the antibiotic biosynthesis regulator AdmX from the rhizobacterium Serratia plymuthica A153 bound to the auxin indole-3-acetic acid (IAA). | | Descriptor: | 1H-INDOL-3-YLACETIC ACID, MAGNESIUM ION, TRANSCRIPTIONAL REGULATOR AdmX | | Authors: | Gavira, J.A, Rico-Jimenez, M, Castellvi, A, Krell, T, Matilla, M.A. | | Deposit date: | 2021-12-03 | | Release date: | 2022-12-14 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Emergence of an Auxin Sensing Domain in Plant-Associated Bacteria.
Mbio, 14, 2023
|
|
7QEK
 | | Structure of the ligand binding domain of the antibiotic biosynthesis regulator AdmX from the rhizobacterium Serratia plymuthica A153 bound to the auxin indole-3-piruvic acid (IPA). | | Descriptor: | 3-(1H-INDOL-3-YL)-2-OXOPROPANOIC ACID, MAGNESIUM ION, regulator AdmX | | Authors: | Gavira, J.A, Rico-Jimenez, M, Castellvi, A, Krell, T, Matilla, M.A. | | Deposit date: | 2021-12-03 | | Release date: | 2022-12-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Emergence of an Auxin Sensing Domain in Plant-Associated Bacteria.
Mbio, 14, 2023
|
|
7RRH
 
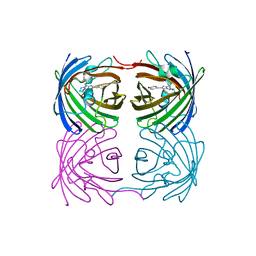 | | Crystal structure of fast switching R66M/M159T mutant of fluorescent protein Dronpa (Dronpa2) | | Descriptor: | Fluorescent protein Dronpa | | Authors: | Lin, C.-Y, Romei, M.G, Mathews, I.I, Boxer, S.G. | | Deposit date: | 2021-08-09 | | Release date: | 2021-10-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.747 Å) | | Cite: | Energetic Basis and Design of Enzyme Function Demonstrated Using GFP, an Excited-State Enzyme.
J.Am.Chem.Soc., 144, 2022
|
|
7RRK
 | | Crystal structure of fast switching M159E mutant of fluorescent protein Dronpa (Dronpa2) | | Descriptor: | Fluorescent protein Dronpa | | Authors: | Lin, C.-Y, Romei, M.G, Mathews, I.I, Boxer, S.G. | | Deposit date: | 2021-08-09 | | Release date: | 2021-10-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.929 Å) | | Cite: | Energetic Basis and Design of Enzyme Function Demonstrated Using GFP, an Excited-State Enzyme.
J.Am.Chem.Soc., 144, 2022
|
|
6MIF
 
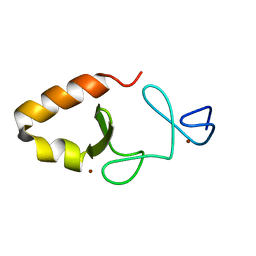 | | Lim5 domain of PINCH1 protein | | Descriptor: | LIM and senescent cell antigen-like-containing domain protein 1, ZINC ION | | Authors: | Qin, J, Vaynberg, J. | | Deposit date: | 2018-09-19 | | Release date: | 2018-10-31 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Non-catalytic signaling by pseudokinase ILK for regulating cell adhesion.
Nat Commun, 9, 2018
|
|
7RRJ
 
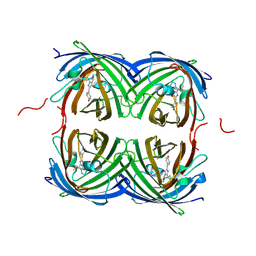 | | Crystal structure of fast switching M159Q mutant of fluorescent protein Dronpa (Dronpa2) | | Descriptor: | Fluorescent protein Dronpa | | Authors: | Lin, C.-Y, Romei, M.G, Mathews, I.I, Boxer, S.G. | | Deposit date: | 2021-08-09 | | Release date: | 2021-10-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Energetic Basis and Design of Enzyme Function Demonstrated Using GFP, an Excited-State Enzyme.
J.Am.Chem.Soc., 144, 2022
|
|
7RRI
 | | Crystal structure of fast switching S142A/M159T mutant of fluorescent protein Dronpa (Dronpa2) | | Descriptor: | Fluorescent protein Dronpa | | Authors: | Lin, C.-Y, Romei, M.G, Mathews, I.I, Boxer, S.G. | | Deposit date: | 2021-08-09 | | Release date: | 2021-10-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.643 Å) | | Cite: | Energetic Basis and Design of Enzyme Function Demonstrated Using GFP, an Excited-State Enzyme.
J.Am.Chem.Soc., 144, 2022
|
|
8DQO
 
 | | Crystal structure of Arabidopsis thaliana COSY | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, Coumarin Synthase | | Authors: | Kim, C.Y, Mitchell, A.J, Gutierrez, M, Weng, J.K. | | Deposit date: | 2022-07-19 | | Release date: | 2023-01-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Emergence of a proton exchange-based isomerization and lactonization mechanism in the plant coumarin synthase COSY.
Nat Commun, 14, 2023
|
|
8DQR
 
 | | Crystal structure of Arabidopsis thaliana COSY in complex with scopoletin | | Descriptor: | CALCIUM ION, COENZYME A, Coumarin Synthase, ... | | Authors: | Kim, C.Y, Mitchell, A.J, Gutierrez, M, Weng, J.K. | | Deposit date: | 2022-07-19 | | Release date: | 2023-01-11 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Emergence of a proton exchange-based isomerization and lactonization mechanism in the plant coumarin synthase COSY.
Nat Commun, 14, 2023
|
|
3MKQ
 
 | |
8DQQ
 
 | | Crystal structure of Arabidopsis thaliana COSY in complex with scopoletin | | Descriptor: | 7-hydroxy-2H-chromen-2-one, ACETATE ION, CALCIUM ION, ... | | Authors: | Kim, C.Y, Mitchell, A.J, Gutierrez, M, Weng, J.K. | | Deposit date: | 2022-07-19 | | Release date: | 2023-01-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Emergence of a proton exchange-based isomerization and lactonization mechanism in the plant coumarin synthase COSY.
Nat Commun, 14, 2023
|
|
8DQP
 
 | | Crystal structure of Arabidopsis thaliana COSY in complex with scopoletin | | Descriptor: | 1,2-ETHANEDIOL, 7-hydroxy-6-methoxy-2H-1-benzopyran-2-one, ACETATE ION, ... | | Authors: | Kim, C.Y, Mitchell, A.J, Gutierrez, M, Weng, J.K. | | Deposit date: | 2022-07-19 | | Release date: | 2023-01-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Emergence of a proton exchange-based isomerization and lactonization mechanism in the plant coumarin synthase COSY.
Nat Commun, 14, 2023
|
|
1ZUN
 
 | | Crystal Structure of a GTP-Regulated ATP Sulfurylase Heterodimer from Pseudomonas syringae | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, ... | | Authors: | Mougous, J.D, Lee, D.H, Hubbard, S.C, Schelle, M.W, Vocadlo, D.J, Berger, J.M, Bertozzi, C.R. | | Deposit date: | 2005-05-31 | | Release date: | 2006-01-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Molecular basis for g protein control of the prokaryotic ATP sulfurylase.
Mol.Cell, 21, 2006
|
|
3PCO
 
 | | crystal structure of E. coli phenylalanine-tRNA synthetase complexed with phenylalanine and AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, PHENYLALANINE, Phenylalanyl-tRNA synthetase, ... | | Authors: | Mermershtain, I, Finarov, I, Klipcan, L, Kessler, N, Rozenberg, H, Safro, M.G. | | Deposit date: | 2010-10-21 | | Release date: | 2011-03-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.02 Å) | | Cite: | Idiosyncrasy and identity in the prokaryotic phe-system: crystal structure of E. coli phenylalanyl-tRNA synthetase complexed with phenylalanine and AMP.
Protein Sci., 20, 2011
|
|
1XFP
 
 | | Crystal structure of the CDR2 germline reversion mutant of cAb-Lys3 in complex with hen egg white lysozyme | | Descriptor: | FORMIC ACID, Lysozyme C, heavy chain antibody | | Authors: | De Genst, E, Handelberg, F, Van Meirhaeghe, A, Vynck, S, Loris, R, Wyns, L, Muyldermans, S. | | Deposit date: | 2004-09-15 | | Release date: | 2004-09-28 | | Last modified: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Chemical Basis for the Affinity Maturation of a Camel Single Domain Antibody
J.Biol.Chem., 279, 2004
|
|
2L40
 
 | | Mouse prion protein (121-231) containing the substitution Y169A | | Descriptor: | Major prion protein | | Authors: | Christen, B, Damberger, F.F, Perez, D.R, Hornemann, S, Wuthrich, K. | | Deposit date: | 2010-09-28 | | Release date: | 2011-08-10 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Cellular prion protein conformation and function.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
2L39
 
 | | Mouse prion protein fragment 121-231 AT 37 C | | Descriptor: | Major prion protein | | Authors: | Christen, B, Damberger, F.F, Perez, D.R, Hornemann, S, Wuthrich, K. | | Deposit date: | 2010-09-10 | | Release date: | 2011-08-10 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Cellular prion protein conformation and function.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
